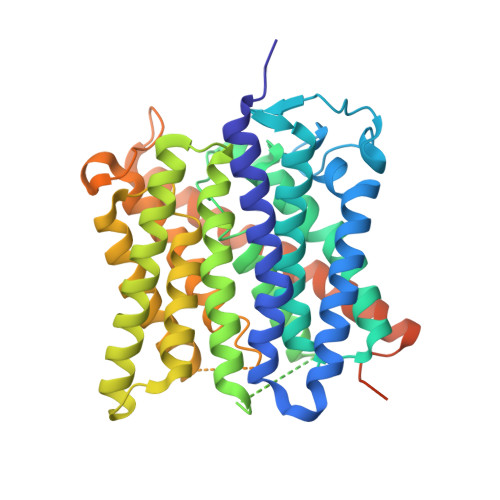
An unusual twin-his arrangement in the pore of ammonia channels is essential for substrate conductance
Javelle, A., Lupo, D., Zheng, L., Li, X.-D., Winkler, F.K., Merrick, M.(2006) J Biological Chem 281: 39492-39498
- PubMed: 17040913 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M608325200
- Primary Citation Related Structures:
2NMR, 2NOP, 2NOW, 2NPC, 2NPD, 2NPE, 2NPG, 2NPJ, 2NPK - PubMed Abstract:
Amt proteins constitute a class of ubiquitous integral membrane proteins that mediate movement of ammonium across cell membranes. They are homotrimers, in which each subunit contains a narrow pore through which substrate transport occurs. Two conserved histidine residues in the pore have been proposed to be necessary for ammonia conductance. By analyzing 14 engineered polar and non-polar variants of these histidines, in Escherichia coli AmtB, we show that both histidines are absolutely required for optimum substrate conductance. Crystal structures of variants confirm that substitution of the histidine residues does not affect AmtB structure. In a subgroup of Amt proteins, found only in fungi, one of the histidines is replaced by glutamate. The equivalent substitution in E. coli AmtB is partially active, and the structure of this variant suggests that the glutamate side chain can make similar interactions to those made by histidine.
- Department of Molecular Microbiology, John Innes Centre, Colney Lane, Norwich, Norfolk, NR4 7UH, United Kingdom.
Organizational Affiliation: