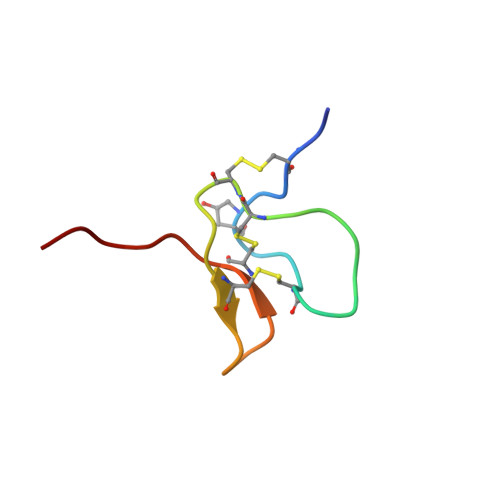
Structural Basis for the Inhibition of Voltage-gated Sodium Channels by Conotoxin mu O-GVIIJ.
Green, B.R., Gajewiak, J., Chhabra, S., Skalicky, J.J., Zhang, M.M., Rivier, J.E., Bulaj, G., Olivera, B.M., Yoshikami, D., Norton, R.S.(2016) J Biological Chem 291: 7205-7220
- PubMed: 26817840 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.697672
- Primary Citation Related Structures:
2N8H - PubMed Abstract:
Cone snail toxins are well known blockers of voltage-gated sodium channels, a property that is of broad interest in biology and therapeutically in treating neuropathic pain and neurological disorders. Although most conotoxin channel blockers function by direct binding to a channel and disrupting its normal ion movement, conotoxin μO§-GVIIJ channel blocking is unique, using both favorable binding interactions with the channel and a direct tether via an intermolecular disulfide bond. Disulfide exchange is possible because conotoxin μO§-GVIIJ contains anS-cysteinylated Cys-24 residue that is capable of exchanging with a free cysteine thiol on the channel surface. Here, we present the solution structure of an analog of μO§-GVIIJ (GVIIJ[C24S]) and the results of structure-activity studies with synthetic μO§-GVIIJ variants. GVIIJ[C24S] adopts an inhibitor cystine knot structure, with two antiparallel β-strands stabilized by three disulfide bridges. The loop region linking the β-strands (loop 4) presents residue 24 in a configuration where it could bind to the proposed free cysteine of the channel (Cys-910, rat NaV1.2 numbering; at site 8). The structure-activity study shows that three residues (Lys-12, Arg-14, and Tyr-16) located in loop 2 and spatially close to residue 24 were also important for functional activity. We propose that the interaction of μO§-GVIIJ with the channel depends on not only disulfide tethering via Cys-24 to a free cysteine at site 8 on the channel but also the participation of key residues of μO§-GVIIJ on a distinct surface of the peptide.
- From the Medicinal Chemistry, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, Victoria 3052, Australia, the Department of Biology, University of Utah, Salt Lake City, Utah 84112.
Organizational Affiliation: