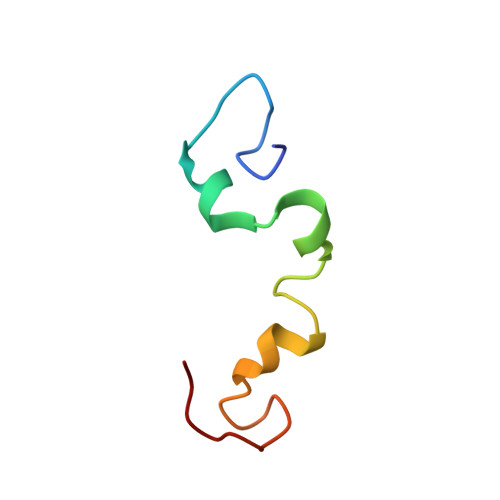
Folding of the Tau Protein on Microtubules.
Kadavath, H., Jaremko, M., Jaremko, M., Biernat, J., Mandelkow, E., Zweckstetter, M.(2015) Angew Chem Int Ed Engl 54: 10347-10351
- PubMed: 26094605 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201501714
- Primary Citation Related Structures:
2MZ7 - PubMed Abstract:
Microtubules are regulated by microtubule-associated proteins. However, little is known about the structure of microtubule-associated proteins in complex with microtubules. Herein we show that the microtubule-associated protein Tau, which is intrinsically disordered in solution, locally folds into a stable structure upon binding to microtubules. While Tau is highly flexible in solution and adopts a β-sheet structure in amyloid fibrils, in complex with microtubules the conserved hexapeptides at the beginning of the Tau repeats two and three convert into a hairpin conformation. Thus, binding to microtubules stabilizes a unique conformation in Tau.
- Max Planck Institute for Biophysical Chemistry, Am Fassberg 11, 37077 Göttingen (Germany).
Organizational Affiliation: