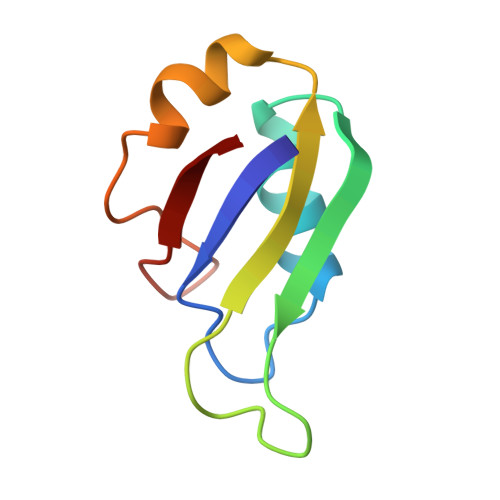
Structure, backbone dynamics and interactions with RNA of the C-terminal RNA-binding domain of a mouse neural RNA-binding protein, Musashi1.
Nagata, T., Kanno, R., Kurihara, Y., Uesugi, S., Imai, T., Sakakibara, S., Okano, H., Katahira, M.(1999) J Mol Biology 287: 315-330
- PubMed: 10080895 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2596
- Primary Citation Related Structures:
2MSS, 2MST - PubMed Abstract:
Musashi1 is an RNA-binding protein abundantly expressed in the developing mouse central nervous system. Its restricted expression in neural precursor cells suggests that it is involved in the regulation of asymmetric cell division. Musashi1 contains two ribonucleoprotein (RNP)-type RNA-binding domains (RBDs), RBD1 and RBD2. Our previous studies showed that RBD1 alone binds to RNA, while the binding of RBD2 is not detected under the same conditions. Joining of RBD2 to RBD1, however, increases the affinity to greater than that of RBD1 alone, indicating that RBD2 contributes to RNA-binding. We have determined the three-dimensional solution structure of the C-terminal RBD (RBD2) of Musashi1 by NMR. It folds into a compact alpha beta structure comprising a four-stranded antiparallel beta-sheet packed against two alpha-helices, which is characteristic of RNP-type RBDs. Special structural features of RBD2 include a beta-bulge in beta2 and a shallow twist of the beta-sheet. The smaller 1H-15N nuclear Overhauser enhancement values for the residues of loop 3 between beta2 and beta3 suggest that this loop is flexible in the time-scale of nano- to picosecond order. The smaller 15N T2 values for the residues around the border between alpha2 and the following loop (loop 5) suggest this region undergoes conformational exchange in the milli- to microsecond time-scale. Chemical shift perturbation analysis indicated that RBD2 binds to an RNA oligomer obtained by in vitro selection under the conditions for NMR measurements, and thus the nature of the weak RNA-binding of RBD2 was successfully characterized by NMR, which is otherwise difficult to assess. Mainly the residues of the surface composed of the four-stranded beta-sheet, loops and C-terminal region are involved in the interaction. The appearance of side-chain NH proton resonances of arginine residues of loop 3 and imino proton resonances of RNA bases upon complex formation suggests the formation of intermolecular hydrogen bonds. The structural arrangement of the rings of the conserved aromatic residues of beta2 and beta3 is suitable for stacking interaction with RNA bases, known to be one of the major protein-RNA interactions, but a survey of the perturbation data suggested that the stacking interaction is not ideally achieved in the complex, which may be related to the weaker RNA-binding of RBD2.
- Department of Chemistry and Biotechnology, Faculty of Engineering, Yokohama National University, 79-5 Tokiwadai, Hodogaya-ku, Yokohama, 240-8501, Japan.
Organizational Affiliation: