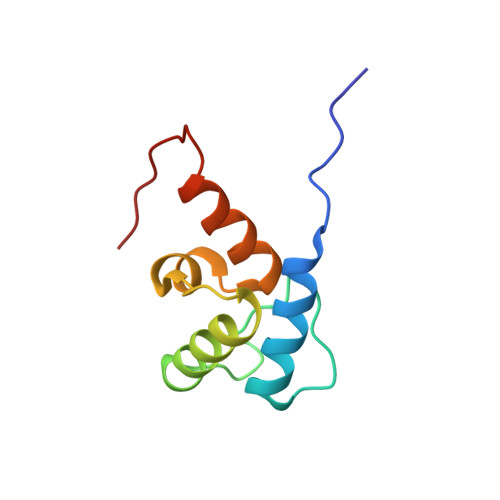
Structure of the terminal PCP domain of the non-ribosomal peptide synthetase in teicoplanin biosynthesis.
Haslinger, K., Redfield, C., Cryle, M.J.(2015) Proteins 83: 711-721
- PubMed: 25586301 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24758
- Primary Citation Related Structures:
2MR7, 2MR8 - PubMed Abstract:
The biosynthesis of the glycopeptide antibiotics, of which teicoplanin and vancomycin are representative members, relies on the combination of non-ribosomal peptide synthesis and modification of the peptide by cytochrome P450 (Oxy) enzymes while the peptide remains bound to the peptide synthesis machinery. We have structurally characterized the final peptidyl carrier protein domain of the teicoplanin non-ribosomal peptide synthetase machinery: this domain is believed to mediate the interactions with tailoring Oxy enzymes in addition to its function as a shuttle for intermediates between multiple non-ribosomal peptide synthetase domains. Using solution state NMR, we have determined structures of this PCP domain in two states, the apo and the post-translationally modified holo state, both of which conform to a four-helix bundle assembly. The structures exhibit the same general fold as the majority of known carrier protein structures, in spite of the complex biosynthetic role that PCP domains from the final non-ribosomal peptide synthetase module must play in glycopeptide antibiotic biosynthesis. These structures thus support the hypothesis that it is subtle rearrangements, rather than dramatic conformational changes, which govern carrier protein interactions and selectivity during non-ribosomal peptide synthesis.
- Department of Biomolecular Mechanisms, Max Planck Institute for Medical Research, Jahnstrasse 29, Heidelberg, 69120, Germany.
Organizational Affiliation: