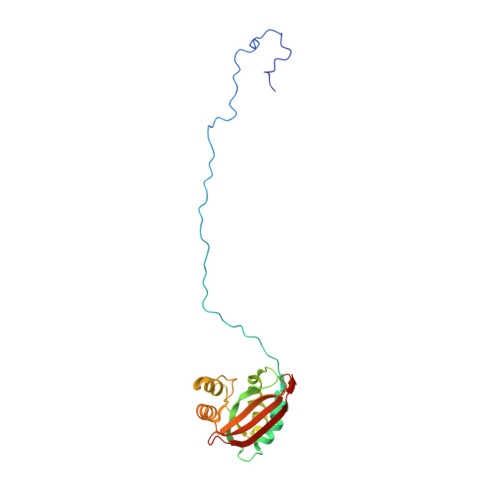
Outer-membrane lipoprotein LpoB spans the periplasm to stimulate the peptidoglycan synthase PBP1B.
Egan, A.J., Jean, N.L., Koumoutsi, A., Bougault, C.M., Biboy, J., Sassine, J., Solovyova, A.S., Breukink, E., Typas, A., Vollmer, W., Simorre, J.P.(2014) Proc Natl Acad Sci U S A 111: 8197-8202
- PubMed: 24821816 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1400376111
- Primary Citation Related Structures:
2MII - PubMed Abstract:
Bacteria surround their cytoplasmic membrane with an essential, stress-bearing peptidoglycan (PG) layer. Growing and dividing cells expand their PG layer by using membrane-anchored PG synthases, which are guided by dynamic cytoskeletal elements. In Escherichia coli, growth of the mainly single-layered PG is also regulated by outer membrane-anchored lipoproteins. The lipoprotein LpoB is required for the activation of penicillin-binding protein (PBP) 1B, which is a major, bifunctional PG synthase with glycan chain polymerizing (glycosyltransferase) and peptide cross-linking (transpeptidase) activities. Here, we report the structure of LpoB, determined by NMR spectroscopy, showing an N-terminal, 54-aa-long flexible stretch followed by a globular domain with similarity to the N-terminal domain of the prevalent periplasmic protein TolB. We have identified the interaction interface between the globular domain of LpoB and the noncatalytic UvrB domain 2 homolog domain of PBP1B and modeled the complex. Amino acid exchanges within this interface weaken the PBP1B-LpoB interaction, decrease the PBP1B stimulation in vitro, and impair its function in vivo. On the contrary, the N-terminal flexible stretch of LpoB is required to stimulate PBP1B in vivo, but is dispensable in vitro. This supports a model in which LpoB spans the periplasm to interact with PBP1B and stimulate PG synthesis.
- Centre for Bacterial Cell Biology.
Organizational Affiliation: