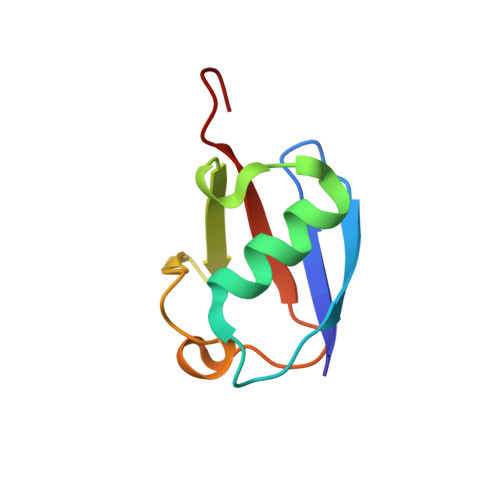
Solution structure of lysine-free (K0) ubiquitin.
Huang, T., Li, J., Byrd, R.A.(2014) Protein Sci 23: 662-667
- PubMed: 24591328 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2450
- Primary Citation Related Structures:
2MI8 - PubMed Abstract:
Lysine-free ubiquitin (K0-Ub) is commonly used to study the ubiquitin-signaling pathway, where it is assumed to have the same structure and function as wild-type ubiquitin (wt-Ub). However, the K0-Ub (15) N heteronuclear single quantum correlation NMR spectrum differs significantly from wt-Ub and the melting temperature is depressed by 19°C, raising the question of the structural integrity and equivalence to wt-Ub. The three-dimensional structure of K0-Ub was determined by solution NMR, using chemical shift and residual dipolar coupling data. K0-Ub adopts the same backbone structure as wt-Ub, and all significant chemical shifts can be related to interactions impacted by the K to R mutations.
- Structural Biophysics Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, Maryland, 21702.
Organizational Affiliation: