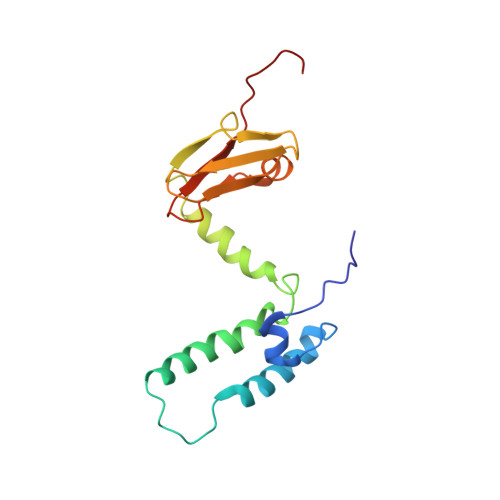
Structure and mechanistic insights into novel iron-mediated moonlighting functions of human J-protein cochaperone, Dph4.
Thakur, A., Chitoor, B., Goswami, A.V., Pareek, G., Atreya, H.S., D'Silva, P.(2012) J Biological Chem 287: 13194-13205
- PubMed: 22367199 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.339655
- Primary Citation Related Structures:
2L6L - PubMed Abstract:
J-proteins are obligate cochaperones of Hsp70s and stimulate their ATPase activity via the J-domain. Although the functions of J-proteins have been well understood in the context of Hsp70s, their additional co-evolved "physiological functions" are still elusive. We report here the solution structure and mechanism of novel iron-mediated functional roles of human Dph4, a type III J-protein playing a vital role in diphthamide biosynthesis and normal development. The NMR structure of Dph4 reveals two domains: a conserved J-domain and a CSL-domain connected via a flexible linker-helix. The linker-helix modulates the conformational flexibility between the two domains, regulating thereby the protein function. Dph4 exhibits a unique ability to bind iron in tetrahedral coordination geometry through cysteines of its CSL-domain. The oxidized Fe-Dph4 shows characteristic UV-visible and electron paramagnetic resonance spectral properties similar to rubredoxins. Iron-bound Dph4 (Fe-Dph4) also undergoes oligomerization, thus potentially functioning as a transient "iron storage protein," thereby regulating the intracellular iron homeostasis. Remarkably, Fe-Dph4 exhibits vital redox and electron carrier activity, which is critical for important metabolic reactions, including diphthamide biosynthesis. Further, we observed that Fe-Dph4 is conformationally better poised to perform Hsp70-dependent functions, thus underlining the significance of iron binding in Dph4. Yeast Jjj3, a functional ortholog of human Dph4 also shows a similar iron-binding property, indicating the conserved nature of iron sequestration across species. Taken together, our findings provide invaluable evidence in favor of additional co-evolved specialized functions of J-proteins, previously not well appreciated.
- Department of Biochemistry, Indian Institute of Science, Bangalore 560012, India.
Organizational Affiliation: