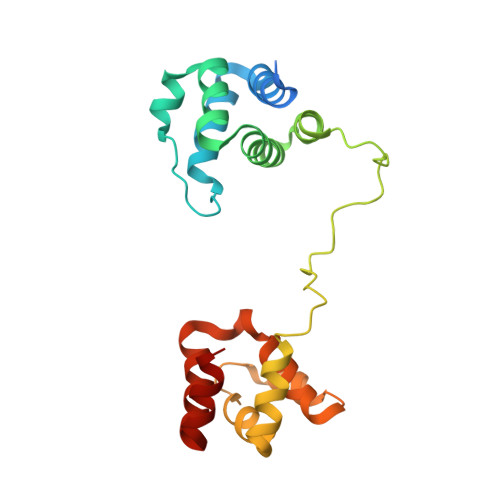
Structure and interdomain dynamics of apoptosis-associated speck-like protein containing a CARD (ASC)
de Alba, E.(2009) J Biological Chem 284: 32932-32941
- PubMed: 19759015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.024273
- Primary Citation Related Structures:
2KN6 - PubMed Abstract:
The human protein ASC is a key mediator in apoptosis and inflammation. Through its two death domains (pyrin and CARD) ASC interacts with cell death executioners, acts as an essential adapter for inflammasome integrity, and oligomerizes into functional supramolecular assemblies. However, these functions are not understood at the structural-dynamic level. This study reports the solution structure and interdomain dynamics of full-length ASC. The pyrin and CARD domains are structurally independent six-helix bundle motifs connected by a 23-residue linker. The CARD structure reveals two distinctive characteristics; helix 1 is not fragmented as in all other known CARDs, and its electrostatic surface shows a uniform distribution of positive and negative charges, whereas these are commonly separated into two areas in other death domains. The linker adopts residual structure resulting in a back-to-back orientation of the domains, which avoids steric interference of each domain with the binding site of the other. NMR relaxation experiments show that the linker is flexible despite the residual structure. This flexibility could help expand the relative volume occupied by each domain, thus increasing the capture radius for effectors. Based on the ASC structure, a tentative model is proposed to illustrate how ASC oligomerizes via CARD and pyrin homophilic interactions. Moreover, ASC oligomers have been analyzed by atomic force microscopy, showing a predominant species of disk-like particles of approximately 12-nm diameter and approximately 1-nm height. Taken together, these results provide structural insight into the behavior of ASC as an adapter molecule.
- Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Científicas, Ramiro de Maeztu 9, Madrid 28040, Spain. dealbae@cib.csic.es
Organizational Affiliation: