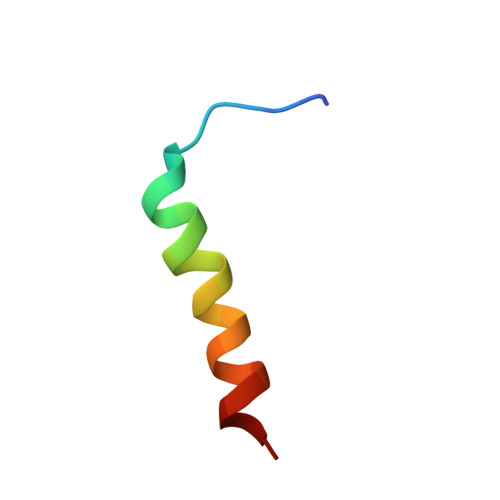
NMR structures of the histidine-rich peptide LAH4 in micellar environments: membrane insertion, pH-dependent mode of antimicrobial action, and DNA transfection.
Georgescu, J., Munhoz, V.H., Bechinger, B.(2010) Biophys J 99: 2507-2515
- PubMed: 20959091
- DOI: https://doi.org/10.1016/j.bpj.2010.05.038
- Primary Citation Related Structures:
2KJN, 2KJO - PubMed Abstract:
The LAH4 family of histidine-rich peptides exhibits potent antimicrobial and DNA transfection activities, both of which require interactions with cellular membranes. The bilayer association of the peptides has been shown to be strongly pH-dependent, with in-planar alignments under acidic conditions and transmembrane orientations when the histidines are discharged. Therefore, we investigated the pH- and temperature-dependent conformations of LAH4 in DPC micellar solutions and in a TFE/PBS solvent mixture. In the presence of detergent and at pH 4.1, LAH4 adopts helical conformations between residues 9 and 24 concomitantly with a high hydrophobic moment. At pH 6.1, a helix-loop-helix structure forms with a hinge encompassing residues His¹⁰-Ala¹³. The data suggest that the high density of histidine residues and the resulting electrostatic repulsion lead to both a decrease in the pK values of the histidines and a less stable α-helical conformation of this region. The hinged structure at pH 6.1 facilitates membrane anchoring and insertion. At pH 7.8, the histidines are uncharged and an extended helical conformation including residues 4-21 is again obtained. LAH4 thus exhibits a high degree of conformational plasticity. The structures provide a stroboscopic view of the conformational changes that occur during membrane insertion, and are discussed in the context of antimicrobial activity and DNA transfection.
- Institut de Chimie, Université de Strasbourg/Centre National de Recherche Scientifique, France.
Organizational Affiliation: