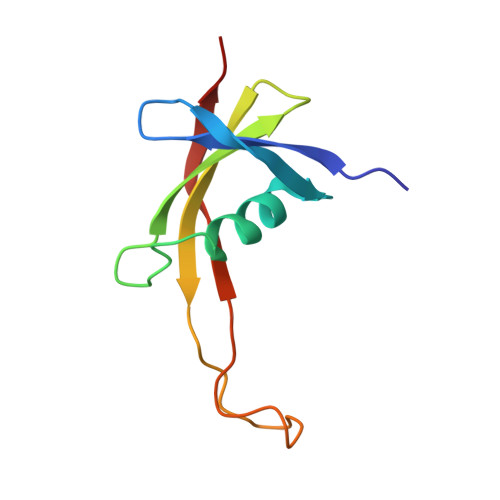
Refined solution structure and backbone dynamics of the archaeal MC1 protein
Paquet, F., Loth, K., Meudal, H., Culard, F., Genest, D., Lancelot, G.(2010) FEBS J 277: 5133-5145
- PubMed: 21078128 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07927.x
- Primary Citation Related Structures:
2KHL - PubMed Abstract:
The 3D structure of methanogen chromosomal protein 1 (MC1), determined with heteronuclear NMR methods, agrees with its function in terms of the shape and nature of the binding surface, whereas the 3D structure determined with homonuclear NMR does not. The structure features five loops, which show a large distribution in the ensemble of 3D structures. Evidence for the fact that this distribution signifies internal mobility on the nanosecond time scale was provided by using (15)N-relaxation and molecular dynamics simulations. Structural variations of the arm (11 residues) induced large shape anisotropy variations on the nanosecond time scale that ruled out the use of the model-free formalism to analyze the relaxation data. The backbone dynamics analysis of MC1 was achieved by comparison with 20 ns molecular dynamics trajectories. Two β-bulges showed that hydrogen bond formation correlated with ϕ and ψ dihedral angle transitions. These jumps were observed on the nanosecond time scale, in agreement with a large decrease in (15)N-NOE for Gly17 and Ile89. One water molecule bridging NH(Glu87) and CO(Val57) through hydrogen bonding contributed to these dynamics. Nanosecond slow motions observed in loops LP3 (35-42) and LP5 (67-77) reflected the lack of stable hydrogen bonds, whereas the other loops, LP1 (10-14), LP2 (22-24), and LP4 (50-53), were stabilized by several hydrogen bonds. Dynamics are often directly related to function. Our data strongly suggest that residues belonging to the flexible regions of MC1 could be involved in the interaction with DNA.
- Centre de Biophysique Moléculaire, CNRS UPR 4301, Orléans, France. francoise.paquet@cnrs-orleans.fr
Organizational Affiliation: