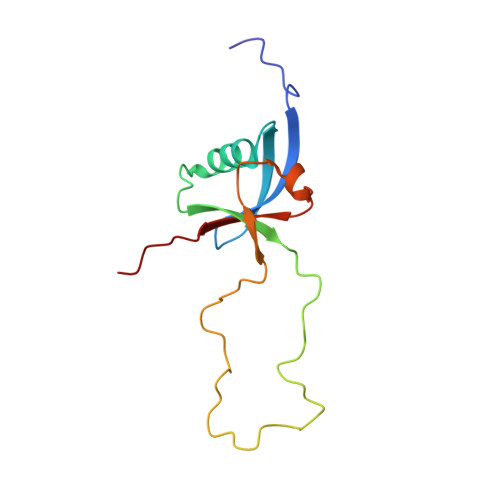
Structure of a double ubiquitin-like domain in the talin head: a role in integrin activation.
Goult, B.T., Bouaouina, M., Elliott, P.R., Bate, N., Patel, B., Gingras, A.R., Grossmann, J.G., Roberts, G.C., Calderwood, D.A., Critchley, D.R., Barsukov, I.L.(2010) EMBO J 29: 1069-1080
- PubMed: 20150896 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2010.4
- Primary Citation Related Structures:
2KC1, 2KC2, 2KMA - PubMed Abstract:
Talin is a 270-kDa protein that activates integrins and couples them to cytoskeletal actin. Talin contains an N-terminal FERM domain comprised of F1, F2 and F3 domains, but it is atypical in that F1 contains a large insert and is preceded by an extra domain F0. Although F3 contains the binding site for beta-integrin tails, F0 and F1 are also required for activation of beta1-integrins. Here, we report the solution structures of F0, F1 and of the F0F1 double domain. Both F0 and F1 have ubiquitin-like folds joined in a novel fixed orientation by an extensive charged interface. The F1 insert forms a loop with helical propensity, and basic residues predicted to reside on one surface of the helix are required for binding to acidic phospholipids and for talin-mediated activation of beta1-integrins. This and the fact that basic residues on F2 and F3 are also essential for integrin activation suggest that extensive interactions between the talin FERM domain and acidic membrane phospholipids are required to orientate the FERM domain such that it can activate integrins.
- Department of Biochemistry, University of Leicester, Leicester, UK.
Organizational Affiliation: