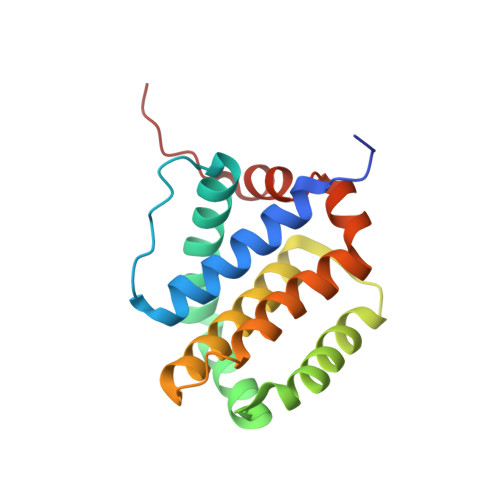
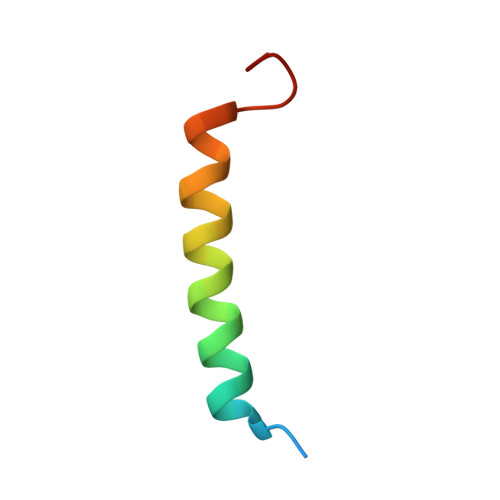
Apoptotic regulation by MCL-1 through heterodimerization.
Liu, Q., Moldoveanu, T., Sprules, T., Matta-Camacho, E., Mansur-Azzam, N., Gehring, K.(2010) J Biological Chem 285: 19615-19624
- PubMed: 20392693 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.105452
- Primary Citation Related Structures:
2KBW - PubMed Abstract:
Myeloid cell leukemia 1 (MCL-1), an anti-apoptotic BCL-2 family member active in the preservation of mitochondrial integrity during apoptosis, has fundamental roles in development and hematopoiesis and is dysregulated in human cancers. It bears a unique, intrinsically unstructured, N-terminal sequence, which leads to its instability in cells and hinders protein production and structural characterization. Here, we present collective data from NMR spectroscopy and titration calorimetry to reveal the selectivity of MCL-1 in binding BCL-2 homology 3 (BH3) ligands of interest for mammalian biology. The N-terminal sequence weakens the BH3 interactions but does not affect selectivity. Its removal by calpain-mediated limited proteolysis results in a stable BCL-2-like core domain of MCL-1 (cMCL-1). This core is necessary and sufficient for BH3 ligand binding. Significantly, we also characterized the in vitro protein-protein interaction between cMCL-1 and activated BID by size exclusion chromatography and NMR titrations. This interaction occurs in a very slow manner in solution but is otherwise similar to the interaction between cMCL-1 and BID-BH3 peptides. We also present the solution structure of complex cMCL-1xhBID-BH3, which completes the family portrait of MCL-1 complexes and may facilitate drug discovery against human tumors.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: