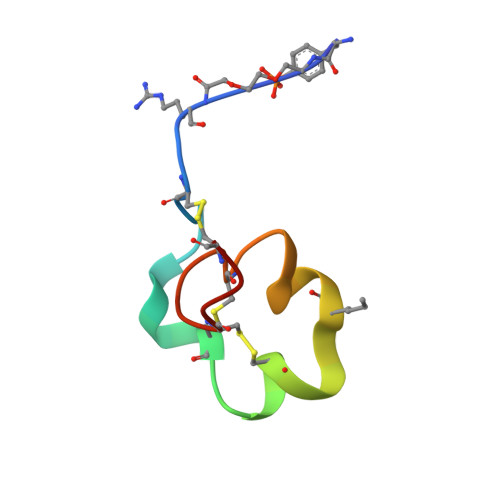
Engineering a stable and selective peptide blocker of the Kv1.3 channel in T lymphocytes
Pennington, M.W., Beeton, C., Galea, C.A., Smith, B.J., Chi, V., Monaghan, K.P., Garcia, A., Rangaraju, S., Giuffrida, A., Plank, D., Crossley, G., Nugent, D., Khaytin, I., Lefievre, Y., Peshenko, I., Dixon, C., Chauhan, S., Orzel, A., Inoue, T., Hu, X., Moore, R.V., Norton, R.S., Chandy, K.G.(2009) Mol Pharmacol 75: 762-773
- PubMed: 19122005 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1124/mol.108.052704
- Primary Citation Related Structures:
2K9E - PubMed Abstract:
Kv1.3 potassium channels maintain the membrane potential of effector memory (T(EM)) T cells that are important mediators of multiple sclerosis, type 1 diabetes mellitus, and rheumatoid arthritis. The polypeptide ShK-170 (ShK-L5), containing an N-terminal phosphotyrosine extension of the Stichodactyla helianthus ShK toxin, is a potent and selective blocker of these channels. However, a stability study of ShK-170 showed minor pH-related hydrolysis and oxidation byproducts that were exacerbated by increasing temperatures. We therefore engineered a series of analogs to minimize the formation of these byproducts. The analog with the greatest stability, ShK-192, contains a nonhydrolyzable phosphotyrosine surrogate, a methionine isostere, and a C-terminal amide. ShK-192 shows the same overall fold as ShK, and there is no evidence of any interaction between the N-terminal adduct and the rest of the peptide. The docking configuration of ShK-192 in Kv1.3 shows the N-terminal para-phosphonophenylalanine group lying at the junction of two channel monomers to form a salt bridge with Lys(411) of the channel. ShK-192 blocks Kv1.3 with an IC(50) of 140 pM and exhibits greater than 100-fold selectivity over closely related channels. After a single subcutaneous injection of 100 microg/kg, approximately 100 to 200 pM concentrations of active peptide is detectable in the blood of Lewis rats 24, 48, and 72 h after the injection. ShK-192 effectively inhibits the proliferation of T(EM) cells and suppresses delayed type hypersensitivity when administered at 10 or 100 microg/kg by subcutaneous injection once daily. ShK-192 has potential as a therapeutic for autoimmune diseases mediated by T(EM) cells.
- Bachem Bioscience Inc., King of Prussia, Pennsylvania, USA.
Organizational Affiliation: