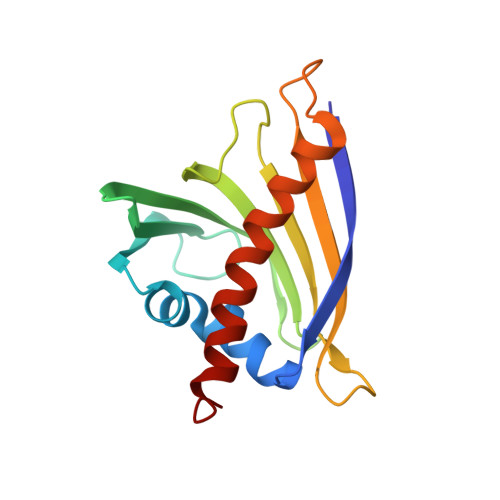
Cross-reactivity of pollen and food allergens: soybean Gly m 4 is a member of the Bet v 1 superfamily and closely resembles yellow lupine proteins
Berkner, H., Neudecker, P., Mittag, D., Ballmer-Weber, B.K., Schweimer, K., Vieths, S.(2009) Biosci Rep 29: 183-192
- PubMed: 18834331 Search on PubMed
- DOI: https://doi.org/10.1042/BSR20080117
- Primary Citation Related Structures:
2K7H - PubMed Abstract:
In many cases, patients allergic to birch pollen also show allergic reactions after ingestion of certain fruits or vegetables. This observation is explained at the molecular level by cross-reactivity of IgE antibodies induced by sensitization to the major birch pollen allergen Bet v 1 with homologous food allergens. As IgE antibodies recognize conformational epitopes, a precise structural characterization of the allergens involved is necessary to understand cross-reactivity and thus to develop new methods of allergen-specific immunotherapy for allergic patients. Here, we report the three-dimensional solution structure of the soybean allergen Gly m 4, a member of the superfamily of Bet v 1 homologous proteins and a cross-reactant with IgE antibodies originally raised against Bet v 1 as shown by immunoblot inhibition and histamine release assays. Although the overall fold of Gly m 4 is very similar to that of Bet v 1, the three-dimensional structures of these proteins differ in detail. The Gly m 4 local structures that display those differences are also found in proteins from yellow lupine with known physiological function. The three-dimensional structure of Gly m 4 may thus shed some light on the physiological function of this subgroup of PR10 proteins (class 10 of pathogenesis-related proteins) and, in combination with immunological data, allow us to propose surface patches that might represent cross-reactive epitopes.
- Department of Biopolymers and Research Center for Bio-Macromolecules, Universität Bayreuth, Universitätsstrasse 30, 95447 Bayreuth, Germany.
Organizational Affiliation: