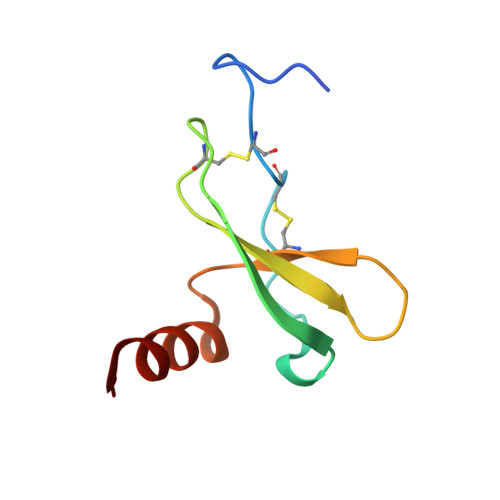
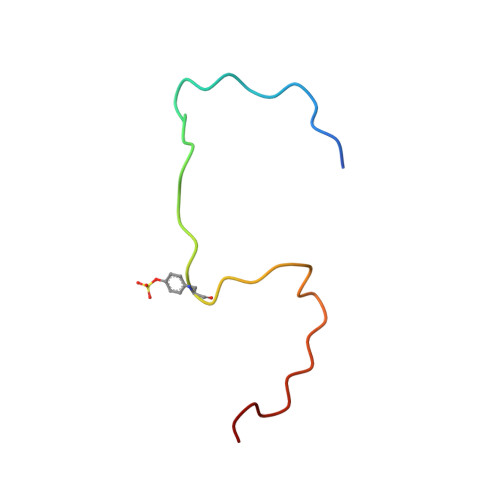
Structural basis of CXCR4 sulfotyrosine recognition by the chemokine SDF-1/CXCL12
Veldkamp, C.T., Seibert, C., Peterson, F.C., De la Cruz, N.B., Haugner, J.C., Basnet, H., Sakmar, T.P., Volkman, B.F.(2008) Sci Signal 1: ra4-ra4
- PubMed: 18799424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scisignal.1160755
- Primary Citation Related Structures:
2K01, 2K03, 2K04, 2K05 - PubMed Abstract:
Stem cell homing and breast cancer metastasis are orchestrated by the chemokine stromal cell-derived factor 1 (SDF-1) and its receptor CXCR4. Here, we report the nuclear magnetic resonance structure of a constitutively dimeric SDF-1 in complex with a CXCR4 fragment that contains three sulfotyrosine residues important for a high-affinity ligand-receptor interaction. CXCR4 bridged the SDF-1 dimer interface so that sulfotyrosines sTyr7 and sTyr12 of CXCR4 occupied positively charged clefts on opposing chemokine subunits. Dimeric SDF-1 induced intracellular Ca2+ mobilization but had no chemotactic activity; instead, it prevented native SDF-1-induced chemotaxis, suggesting that it acted as a potent partial agonist. Our work elucidates the structural basis for sulfotyrosine recognition in the chemokine-receptor interaction and suggests a strategy for CXCR4-targeted drug development.
- Department of Biochemistry, Medical College of Wisconsin, Milwaukee, WI 53226, USA.
Organizational Affiliation: