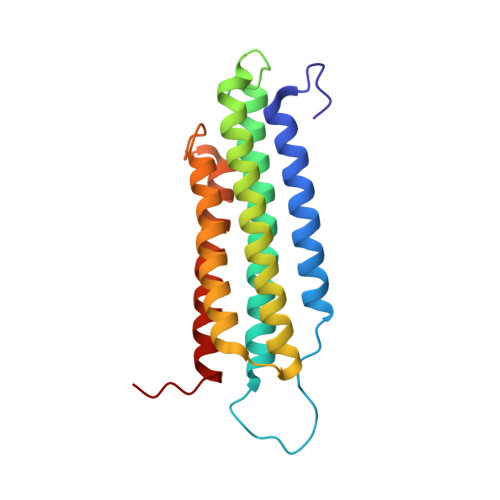
The structure of the C-terminal actin-binding domain of talin.
Gingras, A.R., Bate, N., Goult, B.T., Hazelwood, L., Canestrelli, I., Grossmann, J.G., Liu, H., Putz, N.S., Roberts, G.C., Volkmann, N., Hanein, D., Barsukov, I.L., Critchley, D.R.(2007) EMBO J 27: 458-469
- PubMed: 18157087 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601965
- Primary Citation Related Structures:
2JSW, 2QDQ - PubMed Abstract:
Talin is a large dimeric protein that couples integrins to cytoskeletal actin. Here, we report the structure of the C-terminal actin-binding domain of talin, the core of which is a five-helix bundle linked to a C-terminal helix responsible for dimerisation. The NMR structure of the bundle reveals a conserved surface-exposed hydrophobic patch surrounded by positively charged groups. We have mapped the actin-binding site to this surface and shown that helix 1 on the opposite side of the bundle negatively regulates actin binding. The crystal structure of the dimerisation helix reveals an antiparallel coiled-coil with conserved residues clustered on the solvent-exposed face. Mutagenesis shows that dimerisation is essential for filamentous actin (F-actin) binding and indicates that the dimerisation helix itself contributes to binding. We have used these structures together with small angle X-ray scattering to derive a model of the entire domain. Electron microscopy provides direct evidence for binding of the dimer to F-actin and indicates that it binds to three monomers along the long-pitch helix of the actin filament.
- Department of Biochemistry, University of Leicester, Leicester, UK.
Organizational Affiliation: