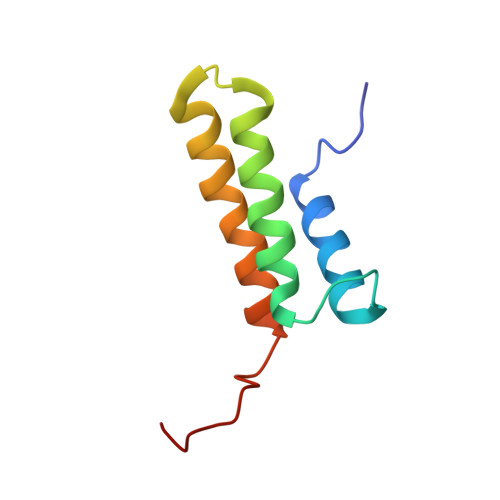
NMR Structure of the Bacillus subtilis Protein YvfG.
Macnaughtan, M.A., Weldeghiorghis, T., Wang, X., Bansal, S., Tian, F., Wang, D., Janjua, H., Cunningham, K., Ma, L.-C., Xiao, R., Liu, J., Baran, M.C., Swapna, G.V.T., Acton, T.B., Rost, B., Montelione, G.T., Prestegard, J.H.To be published.