Exploring Tar-RNA Aptamer Loop-Loop Interaction by X-Ray Crystallography, Uv Spectroscopy and Surface Plasmon Resonance.
Lebars, I., Legrand, P., Aime, A., Pinaud, N., Fribourg, S., Di Primo, C.(2008) Nucleic Acids Res 36: 7146
- PubMed: 18996893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkn831
- Primary Citation Related Structures:
2JLT - PubMed Abstract:
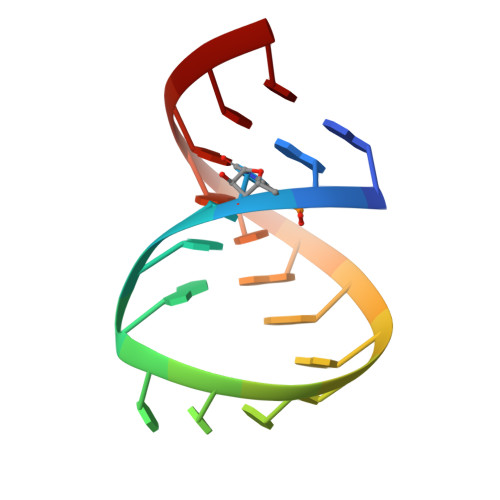
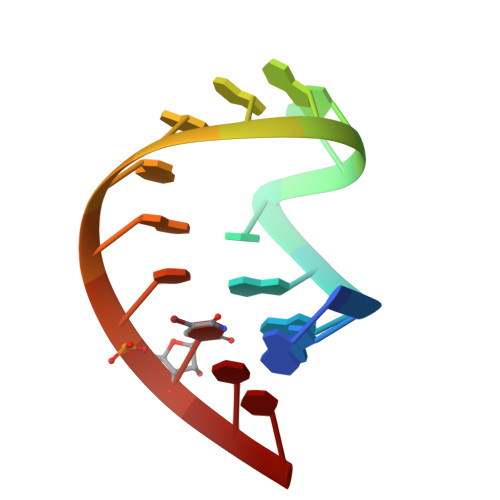
In HIV-1, trans-activation of transcription of the viral genome is regulated by an imperfect hairpin, the trans-activating responsive (TAR) RNA element, located at the 5' untranslated end of all viral transcripts. TAR acts as a binding site for viral and cellular proteins. In an attempt to identify RNA ligands that would interfere with the virus life-cycle by interacting with TAR, an in vitro selection was previously carried out. RNA hairpins that formed kissing-loop dimers with TAR were selected [Ducongé F. and Toulmé JJ (1999) RNA, 5:1605-1614]. We describe here the crystal structure of TAR bound to a high-affinity RNA aptamer. The two hairpins form a kissing complex and interact through six Watson-Crick base pairs. The complex adopts an overall conformation with an inter-helix angle of 28.1 degrees , thus contrasting with previously reported solution and modelling studies. Structural analysis reveals that inter-backbone hydrogen bonds between ribose 2' hydroxyl and phosphate oxygens at the stem-loop junctions can be formed. Thermal denaturation and surface plasmon resonance experiments with chemically modified 2'-O-methyl incorporated into both hairpins at key positions, clearly demonstrate the involvement of this intermolecular network of hydrogen bonds in complex stability.
- CNRS-Université Bordeaux 1-ENITAB, UMR 5248 CBMN, Institut Européen de Chimie et Biologie, Pessac, France.
Organizational Affiliation: