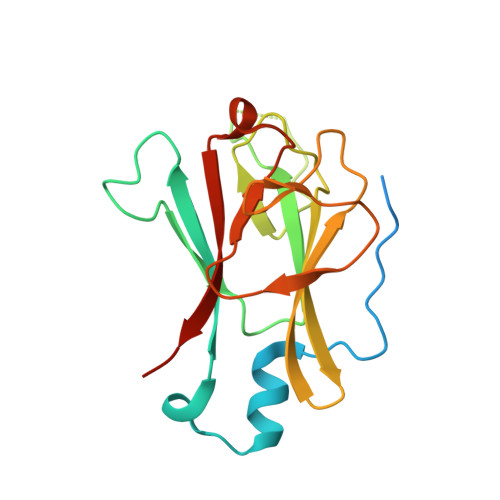
Structure of the Yeast Pml1 Splicing Factor and its Integration Into the Res Complex
Brooks, M.A., Dziembowski, A., Quevillon-Cheruel, S., Henriot, V., Faux, C., Van Tilbeurgh, H., Seraphin, B.(2009) Nucleic Acids Res 37: 129
- PubMed: 19033360 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkn894
- Primary Citation Related Structures:
2JKD - PubMed Abstract:
The RES complex was previously identified in yeast as a splicing factor affecting nuclear pre-mRNA retention. This complex was shown to contain three subunits, namely Snu17, Bud13 and Pml1, but its mode of action remains ill-defined. To obtain insights into its function, we have performed a structural investigation of this factor. Production of a short N-terminal truncation of residues that are apparently disordered allowed us to determine the X-ray crystallographic structure of Pml1. This demonstrated that it consists mainly of a FHA domain, a fold which has been shown to mediate interactions with phosphothreonine-containing peptides. Using a new sensitive assay based on alternative splice-site choice, we show, however, that mutation of the putative phosphothreonine-binding pocket of Pml1 does not affect pre-mRNA splicing. We have also investigated how Pml1 integrates into the RES complex. Production of recombinant complexes, combined with serial truncation and mutagenesis of their subunits, indicated that Pml1 binds to Snu17, which itself contacts Bud13. This analysis allowed us to demarcate the binding sites involved in the formation of this assembly. We propose a model of the organization of the RES complex based on these results, and discuss the functional consequences of this architecture.
- IBBMC-CNRS UMR8619, IFR 115, Université Paris-Sud, Orsay, France.
Organizational Affiliation: