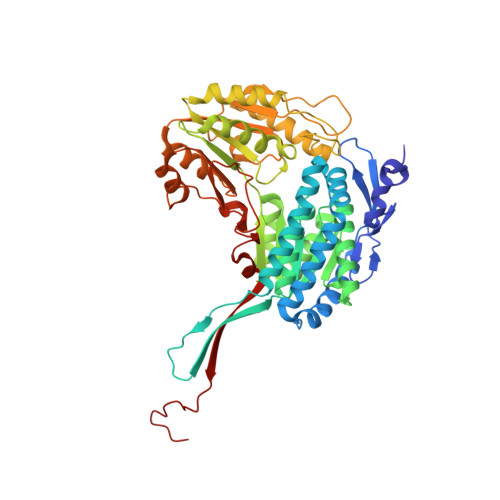
The Crystal Structure of Seabream Antiquitin Reveals the Structural Basis of its Substrate Specificity.
Tang, W.K., Wong, K.B., Lam, Y.M., Cha, S.S., Cheng, C.H.K., Fong, W.P.(2008) FEBS Lett 582: 3090
- PubMed: 18694748
- DOI: https://doi.org/10.1016/j.febslet.2008.07.059
- Primary Citation Related Structures:
2JG7 - PubMed Abstract:
The crystal structure of seabream antiquitin in complex with the cofactor NAD(+) was solved at 2.8A resolution. The mouth of the substrate-binding pocket is guarded by two conserved residues, Glu120 and Arg300. To test the role of these two residues, we have prepared the two mutants E120A and R300A. Our model and kinetics data suggest that antiquitin's specificity towards the substrate alpha-aminoadipic semialdehyde is contributed mainly by Glu120 which interacts with the alpha-amino group of the substrate. On the other hand, Arg300 does not have any specific interaction with the alpha-carboxylate group of the substrate, but is important in maintaining the active site conformation.
- Department of Biochemistry, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong, China.
Organizational Affiliation: