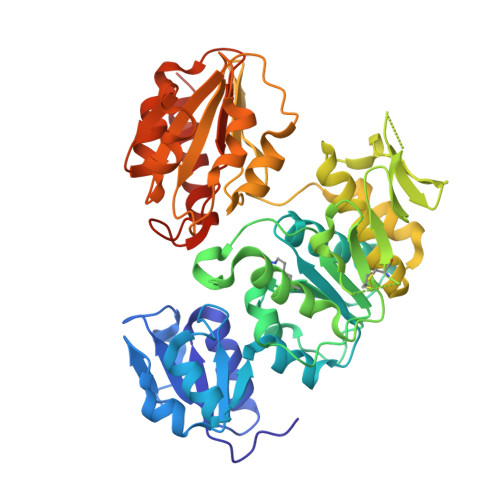
Structural and Functional Characterization of Enantiomeric Glutamic Acid Derivatives as Potential Transition State Analogue Inhibitors of Murd Ligase.
Kotnik, M., Humljan, J., Contreras-Martel, C., Oblak, M., Kristan, K., Herve, M., Blanot, D., Urleb, U., Gobec, S., Dessen, A., Solmajer, T.(2007) J Mol Biology 370: 107
- PubMed: 17507028 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.04.048
- Primary Citation Related Structures:
2JFF, 2JFG, 2JFH - PubMed Abstract:
Mur ligases play an essential role in the intracellular biosynthesis of bacterial peptidoglycan, the main component of the bacterial cell wall, and represent attractive targets for the design of novel antibacterials. UDP-N-acetylmuramoyl-L-alanine:D-glutamate ligase (MurD) catalyses the addition of D-glutamic acid to the cytoplasmic intermediate UDP-N-acetylmuramoyl-L-alanine (UMA) and is the second in the series of Mur ligases. MurD ligase is highly stereospecific for its substrate, D-glutamic acid (D-Glu). Here, we report the high resolution crystal structures of MurD in complexes with two novel inhibitors designed to mimic the transition state of the reaction, which contain either the D-Glu or the L-Glu moiety. The binding modes of N-sulfonyl-D-Glu and N-sulfonyl-L-Glu derivatives were also characterised kinetically. The results of this study represent an excellent starting point for further development of novel inhibitors of this enzyme.
- Lek Pharmaceuticals d. d., Drug Discovery, Verovskova 57, 1526 Ljubljana, Slovenia. miha.kotnik@sandoz.com <miha.kotnik@sandoz.com>
Organizational Affiliation: