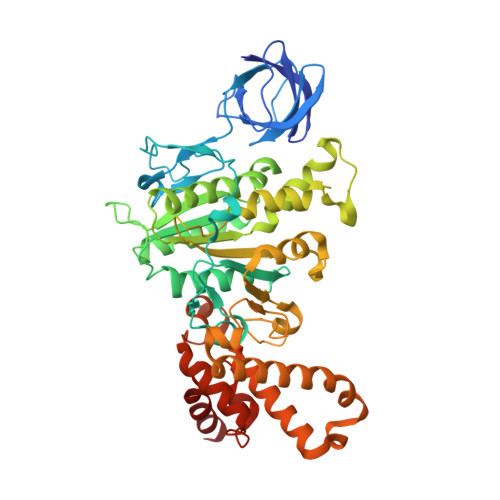
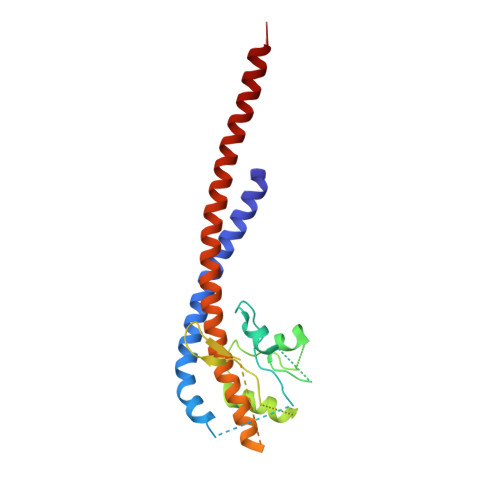
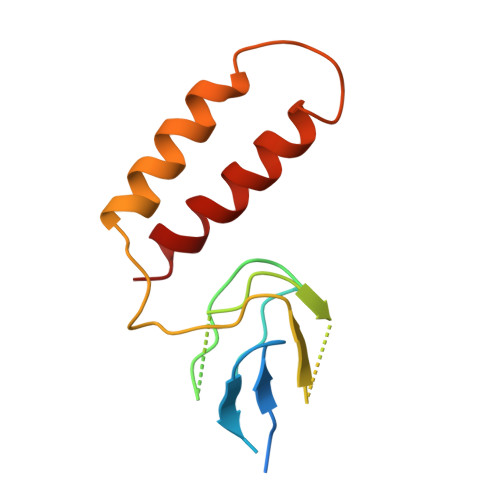
Ground State Structure of F1-ATPase from Bovine Heart Mitochondria at 1.9 A Resolution
Bowler, M.W., Montgomery, M.G., Leslie, A.G.W., Walker, J.E.(2007) J Biological Chem 282: 14238
- PubMed: 17350959 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M700203200
- Primary Citation Related Structures:
2JDI - PubMed Abstract:
The structure of bovine F(1)-ATPase, crystallized in the presence of AMP-PNP and ADP, but in the absence of azide, has been determined at 1.9A resolution. This structure has been compared with the previously described structure of bovine F(1)-ATPase determined at 1.95A resolution with crystals grown under the same conditions but in the presence of azide. The two structures are extremely similar, but they differ in the nucleotides that are bound to the catalytic site in the beta(DP)-subunit. In the present structure, the nucleotide binding sites in the beta(DP)- and beta(TP)-subunits are both occupied by AMP-PNP, whereas in the earlier structure, the beta(TP) site was occupied by AMP-PNP and the beta(DP) site by ADP, where its binding is enhanced by a bound azide ion. Also, the conformation of the side chain of the catalytically important residue, alphaArg-373 differs in the beta(DP)- and beta(TP)-subunits. Thus, the structure with bound azide represents the ADP inhibited state of the enzyme, and the new structure represents a ground state intermediate in the active catalytic cycle of ATP hydrolysis.
- The Medical Research Council Dunn Human Nutrition Unit, Hills Road, Cambridge, UK.
Organizational Affiliation: