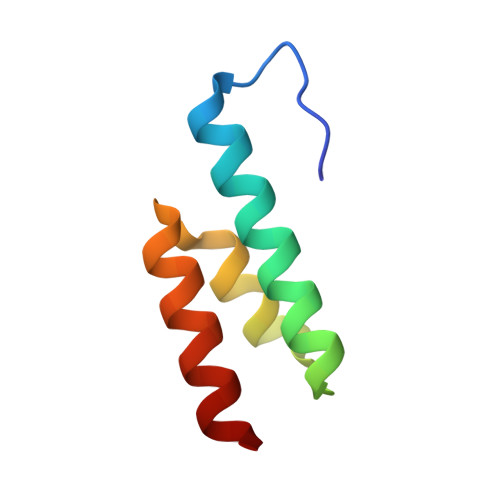
Crystal Structure of a Bacterial Albumin-Binding Domain at 1.4A Resolution.
Cramer, J.F., Nordberg, P.A., Hajdu, J., Lejon, S.(2007) FEBS Lett 581: 3178
- PubMed: 17575979 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.06.003
- Primary Citation Related Structures:
2J5Y - PubMed Abstract:
The albumin-binding domain, or GA module, of the peptostreptococcal albumin-binding protein expressed in pathogenic strains of Finegoldia magna is believed to be responsible for the virulence and increased growth rate of these strains. Here we present the 1.4A crystal structure of this domain, and compare it with the crystal structure of the GA-albumin complex. An analysis of protein-protein interactions in the two crystals, and the presence of multimeric GA species in solution, indicate the GA module is "sticky", and is capable of forming contacts with a range of protein surfaces. This might lead to interactions with different host proteins.
- Department of Cell and Molecular Biology, Uppsala University, Biomedical Centre, Uppsala, Sweden.
Organizational Affiliation: