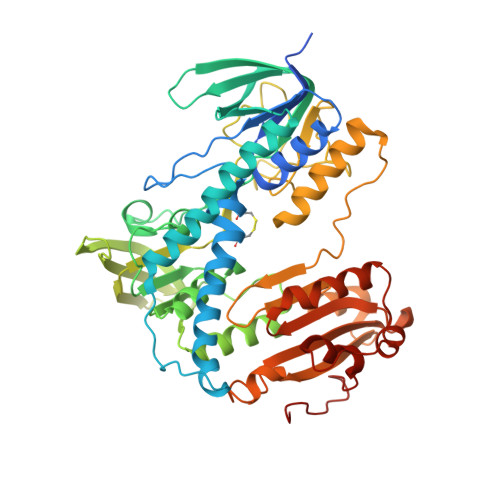
The Structure of Human Thioredoxin Reductase 1 Provides Insights Into C-Terminal Rearrangements During Catalysis.
Fritz-Wolf, K., Urig, S., Becker, K.(2007) J Mol Biology 370: 116
- PubMed: 17512005
- DOI: https://doi.org/10.1016/j.jmb.2007.04.044
- Primary Citation of Related Structures:
2J3N - PubMed Abstract:
Human thioredoxin reductase (hTrxR) is a homodimeric flavoprotein crucially involved in the regulation of cellular redox reactions, growth and differentiation. The enzyme contains a selenocysteine residue at its C-terminal active site that is essential for catalysis. This redox center is located on a flexible arm, solvent-exposed and reactive towards electrophilic inhibitors, thus representing a target for antitumor drug development. During catalysis reducing equivalents are transferred from the cofactor NADPH to FAD, then to the N-terminal active site cysteine residues and from there to the flexible C-terminal part of the other subunit to be finally delivered to a variety of second substrates at the molecule's surface. Here we report the first crystal structure of hTrxR1 (Sec-->Cys) in complex with FAD and NADP(+) at a resolution of 2.8 A. From the crystals three different conformations of the carboxy-terminal arm could be deduced. The predicted movement of the arm is facilitated by the concerted action of the three side-chain residues of N418, N419 and W407, which act as a guiding bar for the C-terminal sliding process. As supported by previous kinetic data, the three visualized conformations might reflect different stages in enzymatic catalysis. Comparison with other disulfide reductases including human glutathione reductase revealed specific inhibitor binding sites in the intersubunit cavity of hTrxR that can be exploited for structure-based inhibitor development.
- Interdisciplinary Research Center, Justus-Liebig-University, 35392 Giessen, Germany.
Organizational Affiliation: