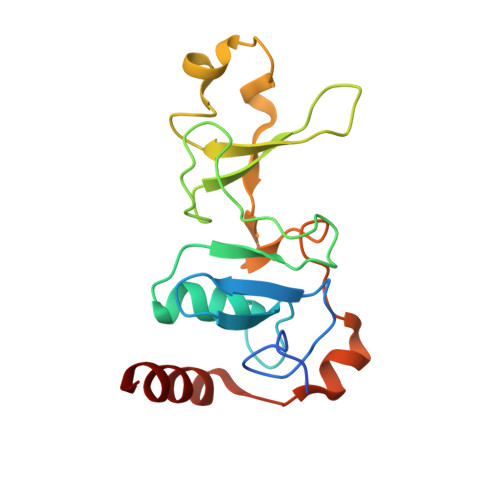
Crystal Structure of the Upf2-Interacting Domain of Nonsense-Mediated Mrna Decay Factor Upf1.
Kadlec, J., Guilligay, D., Ravelli, R.B., Cusack, S.(2006) RNA 12: 1817
- PubMed: 16931876
- DOI: https://doi.org/10.1261/rna.177606
- Primary Citation of Related Structures:
2IYK - PubMed Abstract:
UPF1 is an essential eukaryotic RNA helicase that plays a key role in various mRNA degradation pathways, notably nonsense-mediated mRNA decay (NMD). In combination with UPF2 and UPF3, it forms part of the surveillance complex that detects mRNAs containing premature stop codons and triggers their degradation in all organisms studied from yeast to human. We describe the 3 A resolution crystal structure of the highly conserved cysteine-histidine-rich domain of human UPF1 and show that it is a unique combination of three zinc-binding motifs arranged into two tandem modules related to the RING-box and U-box domains of ubiquitin ligases. This UPF1 domain interacts with UPF2, and we identified by mutational analysis residues in two distinct conserved surface regions of UPF1 that mediate this interaction. UPF1 residues we identify as important for the interaction with UPF2 are not conserved in UPF1 homologs from certain unicellular parasites that also appear to lack UPF2 in their genomes.
- European Molecular Biology Laboratory, Grenoble Outstation, BP 181, 38042 Grenoble Cedex 9, France.
Organizational Affiliation: