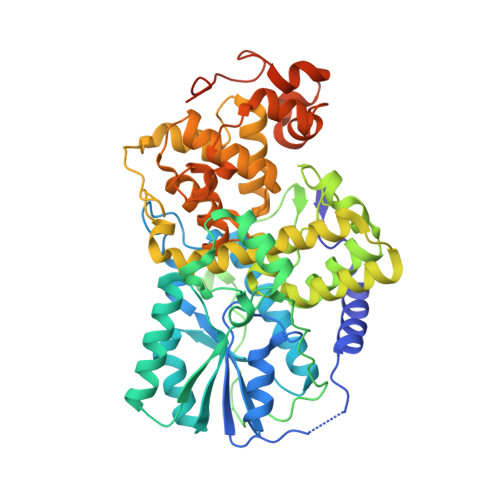
Crystal structure of cryptochrome 3 from Arabidopsis thaliana and its implications for photolyase activity
Huang, Y., Baxter, R., Smith, B.S., Partch, C.L., Colbert, C.L., Deisenhofer, J.(2006) Proc Natl Acad Sci U S A 103: 17701-17706
- PubMed: 17101984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0608554103
- Primary Citation Related Structures:
2IJG - PubMed Abstract:
Cryptochromes use near-UV/blue light to regulate a variety of growth and adaptive process. Recent biochemical studies demonstrate that the Cryptochrome-Drosophila, Arabidopsis, Synechocystis, Human (Cry-DASH) subfamily of cryptochromes have photolyase activity exclusively for single-stranded cyclobutane pyrimidine dimer (CPD)-containing DNA substrate [Selby C, Sancar A (2006) Proc Natl Acad Sci USA 103:17696-17700]. The crystal structure of cryptochrome 3 from Arabidopsis thaliana (At-Cry3), a member of the Cry-DASH proteins, at 2.1 A resolution, reveals that both the light-harvesting cofactor 5,10-methenyl-tetrahydrofolyl-polyglutamate (MTHF) and the catalytic cofactor flavin adenine dinucleotide (FAD) are noncovalently bound to the protein. The residues responsible for binding of MTHF in At-Cry3 are not conserved in Escherichia coli photolyase but are strongly conserved in the Cry-DASH subfamily of cryptochromes. The distance and orientation between MTHF and flavin adenine dinucleotide in At-Cry3 is similar to that of E. coli photolyase, in conjunction with the presence of electron transfer chain, suggesting the conservation of redox activity in At-Cry3. Two amino acid substitutions and the penetration of three charged side chains into the CPD-binding cavity in At-Cry3 alter the hydrophobic environment that is accommodating the hydrophobic sugar ring and thymine base moieties in class I CPD photolyases. These changes most likely make CPD binding less energetically favorable and, hence, insufficient to compete with pairing and stacking interactions between the CPD and the duplex DNA substrate. Thus, Cry-DASH subfamily proteins may be unable to stabilize CPD flipped out from the duplex DNA substrate but may be able to preserve the DNA repair activity toward single-stranded CPD-containing DNA substrate.
- Howard Hughes Medical Institute and Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: