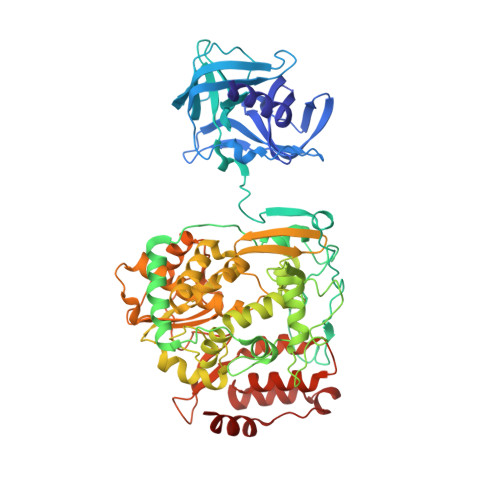
Crystal structure of poliovirus 3CD: virally-encoded protease and precursor to the RNA-dependent RNA polymerase.
Marcotte, L.L., Wass, A.B., Gohara, D.W., Pathak, H.B., Arnold, J.J., Filman, D.J., Cameron, C.E., Hogle, J.M.(2007) J Virol 81: 3583-3596
- PubMed: 17251299 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02306-06
- Primary Citation Related Structures:
2IJD, 2IJF - PubMed Abstract:
Poliovirus 3CD is a multifunctional protein that serves as a precursor to the protease 3C(pro) and the viral polymerase 3D(pol) and also plays a role in the control of viral replication. Although 3CD is a fully functional protease, it lacks polymerase activity. We have solved the crystal structures of 3CD at a 3.4-A resolution and the G64S fidelity mutant of 3D(pol) at a 3.0-A resolution. In the 3CD structure, the 3C and 3D domains are joined by a poorly ordered polypeptide linker, possibly to facilitate its cleavage, in an arrangement that precludes intramolecular proteolysis. The polymerase active site is intact in both the 3CD and the 3D(pol) G64S structures, despite the disruption of a network proposed to position key residues in the active site. Therefore, changes in molecular flexibility may be responsible for the differences in fidelity and polymerase activities. Extensive packing contacts between symmetry-related 3CD molecules and the approach of the 3C domain's N terminus to the VPg binding site suggest how 3D(pol) makes biologically relevant interactions with the 3C, 3CD, and 3BCD proteins that control the uridylylation of VPg during the initiation of viral replication. Indeed, mutations designed to disrupt these interfaces have pronounced effects on the uridylylation reaction in vitro.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115, USA.
Organizational Affiliation: