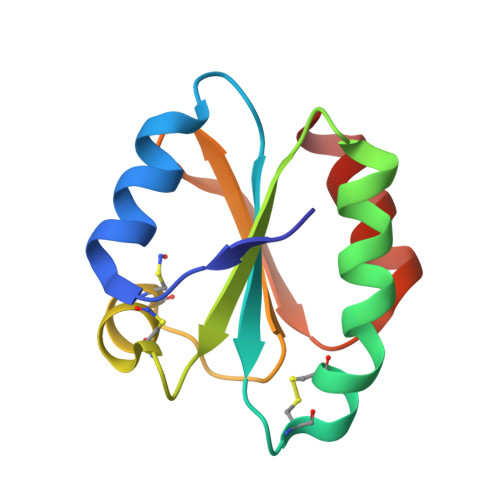
Buried s-nitrosocysteine revealed in crystal structures of human thioredoxin.
Weichsel, A., Brailey, J.L., Montfort, W.R.(2007) Biochemistry 46: 1219-1227
- PubMed: 17260951 Search on PubMed
- DOI: https://doi.org/10.1021/bi061878r
- Primary Citation Related Structures:
2HSH, 2HXK, 2IFQ, 2IIY - PubMed Abstract:
We have determined the 1.65 A crystal structure of human thioredoxin-1 after treatment with S-nitrosoglutathione, providing a high-resolution view of this important protein modification and mechanistic insight into protein transnitrosation. Thioredoxin-1 appears to play an intermediary role in cellular S-nitrosylation and is important in numerous biological and pathobiological activities. S-Nitroso modifications of cysteines 62 and 69 are clearly visible in the structure and display planar cis geometries, whereas cysteines 32, 35, and 73 form intra- and intermolecular disulfide bonds. Surprisingly, the Cys 62 nitroso group is completely buried and pointing to the protein interior yet is the most readily formed at neutral pH. The Cys 69 nitroso group is also protected but requires a higher pH for stable formation. The helix intervening between residues 62 and 69 shifts by approximately 0.5 A to accommodate the SNO groups. The crystallographic asymmetric unit contains three independent molecules of thioredoxin, providing three views of the nitrosated protein. The three molecules are in general agreement but display subtle differences, including both cis and trans conformers for Cys 69 SNO in molecule C, and greater disorder in the Cys 62-Cys 69 helix in molecule B. Possible mechanisms for protein transnitrosation with specific geometric requirements and charge stabilization of the nitroxyl disulfide reaction intermediate are discussed.
- Department of Biochemistry and Molecular Biophysics, University of Arizona, Tucson, Arizona 85721, USA.
Organizational Affiliation: