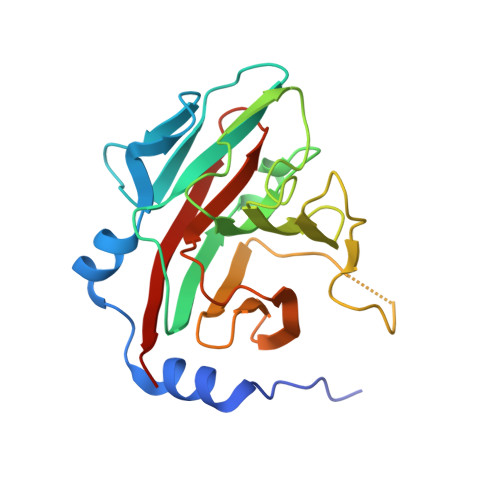
Structural Basis for Protein Recognition by B30.2/SPRY Domains
Woo, J.S., Suh, H.Y., Park, S.Y., Oh, B.H.(2006) Mol Cell 24: 967-976
- PubMed: 17189197 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2006.11.009
- Primary Citation Related Structures:
2IHS - PubMed Abstract:
B30.2/SPRY domains are found in numerous proteins that cover a wide spectrum of biological functions, including regulation of cytokine signaling and innate retroviral restriction. Herein, we report the crystal structure of the B30.2/SPRY domain of a SPRY domain-containing SOCS box (SSB) protein, GUSTAVUS, complexed with a 20 amino acid peptide derived from the RNA helicase VASA, revealing how these domains recognize target proteins. The peptide-binding site is conformationally rigid and has a preformed pocket. The interaction between the pocket and the Asp-Ile-Asn-Asn-Asn-Asn sequence within the peptide accounts for the high-affinity binding between GUSTAVUS and VASA. This observation led to a facile identification of the Glu-Leu-Asn-Asn-Asn-Leu sequence as the recognition motif in a proapoptotic protein Par-4 for its interaction with a GUSTAVUS homolog, SSB-1. Ensuing analyses indicated that many B30.2/SPRY domains have a similar preformed pocket, which would allow them to bind multiple targets.
- Center for Biomolecular Recognition, Department of Life Sciences, Division of Molecular and Life Sciences, Pohang University of Science and Technology, Pohang, Kyungbuk, 790-784, Korea.
Organizational Affiliation: