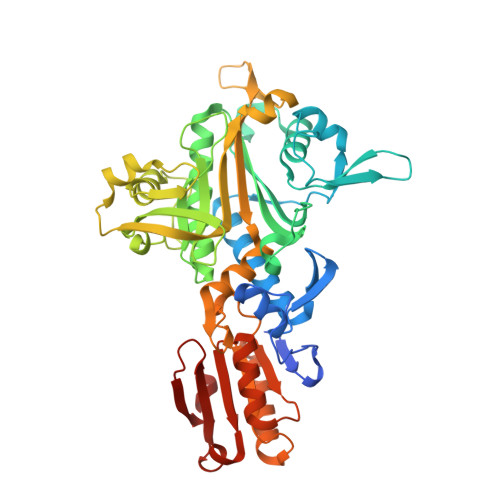
Structures of Two Bacterial Prolyl-tRNA Synthetases with and without a cis-Editing Domain.
Crepin, T., Yaremchuk, A., Tukalo, M., Cusack, S.(2006) Structure 14: 1511-1525
- PubMed: 17027500 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.08.007
- Primary Citation Related Structures:
2I4L, 2I4M, 2I4N, 2I4O, 2J3L, 2J3M - PubMed Abstract:
Prolyl-tRNA synthetases (ProRSs) are unique among synthetases in that they have diverse architectures, notably the variable presence of a cis-editing domain homologous to the freestanding deacylase proteins YbaK and ProX. Here, we describe crystal structures of two bacterial ProRSs from the pathogen Enterococcus faecalis, which possesses an editing domain, and from Rhodopseudomonas palustris, which does not. We compare the overall structure and binding mode of ATP and prolyl-adenylate with those of the archael/eukaryote-type ProRS from Thermus thermophilus. Although structurally more homologous to YbaK, which preferentially hydrolyzes Cys-tRNA(Pro), the editing domain of E. faecalis ProRS possesses key elements similar to ProX, with which it shares the activity of hydrolyzing Ala-tRNA(Pro). The structures give insight into the complex evolution of ProRSs, the mechanism of editing, and structural differences between prokaryotic- and eukaryotic-type ProRSs that can be exploited for antibiotic design.
- European Molecular Biology Laboratory, Grenoble Outstation, 6 rue Jules Horowitz, BP 181, F-38042 Grenoble Cedex 9, France.
Organizational Affiliation: