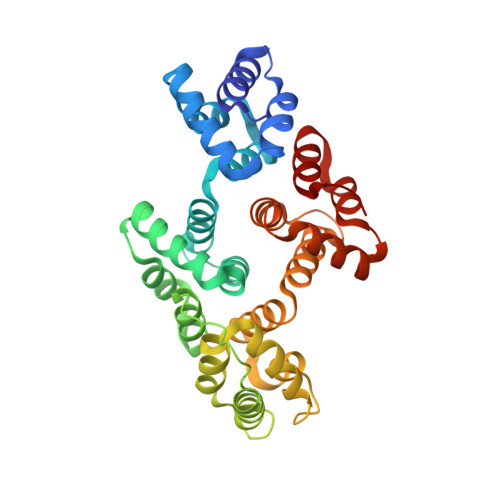
Crystallographic Analysis of Calcium-dependent Heparin Binding to Annexin A2.
Shao, C., Zhang, F., Kemp, M.M., Linhardt, R.J., Waisman, D.M., Head, J.F., Seaton, B.A.(2006) J Biological Chem 281: 31689-31695
- PubMed: 16882661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M604502200
- Primary Citation Related Structures:
2HYU, 2HYV, 2HYW - PubMed Abstract:
Annexin A2 and heparin bind to one another with high affinity and in a calcium-dependent manner, an interaction that may play a role in mediating fibrinolysis. In this study, three heparin-derived oligosaccharides of different lengths were co-crystallized with annexin A2 to elucidate the structural basis of the interaction. Crystal structures were obtained at high resolution for uncomplexed annexin A2 and three complexes of heparin oligosaccharides bound to annexin A2. The common heparin-binding site is situated at the convex face of domain IV of annexin A2. At this site, annexin A2 binds up to five sugar residues from the nonreducing end of the oligosaccharide. Unlike most heparin-binding consensus patterns, heparin binding at this site does not rely on arrays of basic residues; instead, main-chain and side-chain nitrogen atoms and two calcium ions play important roles in the binding. Especially significant is a novel calcium-binding site that forms upon heparin binding. Two sugar residues of the heparin derivatives provide oxygen ligands for this calcium ion. Comparison of all four structures shows that heparin binding does not elicit a significant conformational change in annexin A2. Finally, surface plasmon resonance measurements were made for binding interactions between annexin A2 and heparin polysaccharide in solution at pH 7.4 or 5.0. The combined data provide a clear basis for the calcium dependence of heparin binding to annexin A2.
- Department of Physiology and Biophysics, Boston University School of Medicine, Boston, Massachusetts 02118, USA.
Organizational Affiliation: