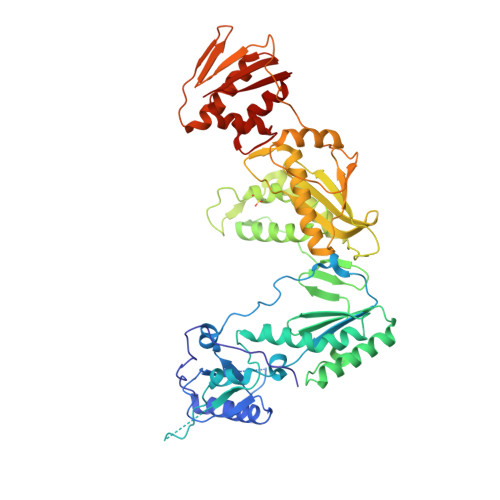
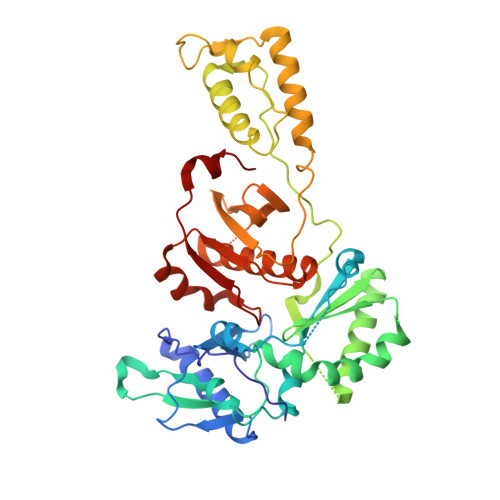
Structural insights into mechanisms of non-nucleoside drug resistance for HIV-1 reverse transcriptases mutated at codons 101 or 138.
Ren, J., Nichols, C.E., Stamp, A., Chamberlain, P.P., Ferris, R., Weaver, K.L., Short, S.A., Stammers, D.K.(2006) FEBS J 273: 3850-3860
- PubMed: 16911530 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05392.x
- Primary Citation Related Structures:
2HND, 2HNY, 2HNZ - PubMed Abstract:
Lys101Glu is a drug resistance mutation in reverse transcriptase clinically observed in HIV-1 from infected patients treated with the non-nucleoside inhibitor (NNRTI) drugs nevirapine and efavirenz. In contrast to many NNRTI resistance mutations, Lys101(p66 subunit) is positioned at the surface of the NNRTI pocket where it interacts across the reverse transcriptase (RT) subunit interface with Glu138(p51 subunit). However, nevirapine contacts Lys101 and Glu138 only indirectly, via water molecules, thus the structural basis of drug resistance induced by Lys101Glu is unclear. We have determined crystal structures of RT(Glu138Lys) and RT(Lys101Glu) in complexes with nevirapine to 2.5 A, allowing the determination of water structure within the NNRTI-binding pocket, essential for an understanding of nevirapine binding. Both RT(Glu138Lys) and RT(Lys101Glu) have remarkably similar protein conformations to wild-type RT, except for significant movement of the mutated side-chains away from the NNRTI pocket induced by charge inversion. There are also small shifts in the position of nevirapine for both mutant structures which may influence ring stacking interactions with Tyr181. However, the reduction in hydrogen bonds in the drug-water-side-chain network resulting from the mutated side-chain movement appears to be the most significant contribution to nevirapine resistance for RT(Lys101Glu). The movement of Glu101 away from the NNRTI pocket can also explain the resistance of RT(Lys101Glu) to efavirenz but in this case is due to a loss of side-chain contacts with the drug. RT(Lys101Glu) is thus a distinctive NNRTI resistance mutant in that it can give rise to both direct and indirect mechanisms of drug resistance, which are inhibitor-dependent.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, Henry Wellcome Building for Genomic Medicine, University of Oxford, UK.
Organizational Affiliation: