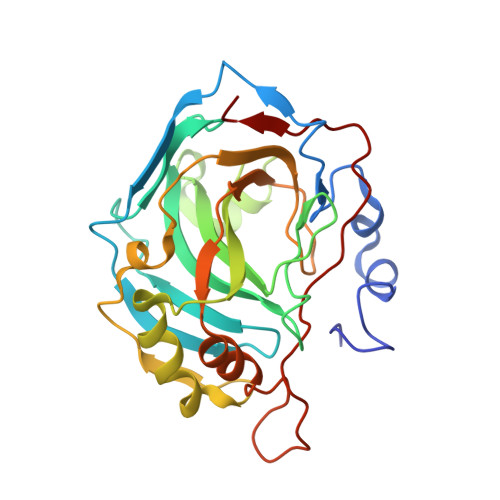
Carbonic anhydrase inhibitors: X-ray crystallographic studies for the binding of 5-amino-1,3,4-thiadiazole-2-sulfonamide and 5-(4-amino-3-chloro-5-fluorophenylsulfonamido)-1,3,4-thiadiazole-2-sulfonamide to human isoform II.
Menchise, V., De Simone, G., Di Fiore, A., Scozzafava, A., Supuran, C.T.(2006) Bioorg Med Chem Lett 16: 6204-6208
- PubMed: 17000110 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.09.022
- Primary Citation Related Structures:
2HNC, 2HOC - PubMed Abstract:
The X-ray crystal structures of 5-amino-1,3,4-thiadiazole-2-sulfonamide (the acetazolamide precursor) and 5-(4-amino-3-chloro-5-fluorophenylsulfonamido)-1,3,4-thiadiazole-2-sulfonamide in complex with the human isozyme II of carbonic anhydrase (CA, EC 4.2.1.1) are reported. The thiadiazole-sulfonamide moiety of the two compounds binds in the canonic manner to the zinc ion and interacts with Thr199, Glu106, and Thr200. The substituted phenyl tail of the second inhibitor was positioned in the hydrophobic part of the binding pocket, at van der Waals distance from Phe131, Val 135, Val141, Leu198, Pro202, and Leu204. These structures may help in the design of better inhibitors of these widespread zinc-containing enzymes.
- Istituto di Biostrutture e Bioimmagini-CNR, via Mezzocannone 16, 80134 Naples, Italy.
Organizational Affiliation: