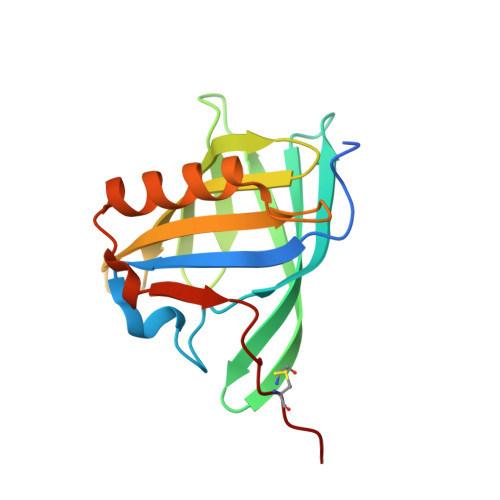
Deswapping bovine odorant binding protein.
Ramoni, R., Spinelli, S., Grolli, S., Conti, V., Merli, E., Cambillau, C., Tegoni, M.(2008) Biochim Biophys Acta 1784: 651-657
- PubMed: 18269920 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2008.01.010
- Primary Citation Related Structures:
2HLV - PubMed Abstract:
The X-ray structure of bovine Odorant Binding Protein (bOBP) revealed its association as a domain swapped dimer. bOBP, devoid of any cysteines, contrasts with other mammalian OBPs, which are monomeric and possess at least one disulfide bridge. We have produced a mutant of bOBP in which a glycine residue was inserted after position 121. This mutation yielded a monomeric bOBP-121Gly+ in which domain swapping has been reverted. Here, we have subsequently introduced two mutations, Trp64Cys and His155Cys, in view to stabilize the putative monomer with a disulfide bridge. We have determined the crystal structure of this triple mutant at 1.65 A resolution. The mutant protein is monomeric, stabilized by a disulfide bridge between Trp64Cys and His155Cys, with a backbone superimposable to that of native bOBP, with the exception of the hinge and of the 10 residues at the C-terminus. bOBP triple mutant binds 1-amino-anthracene, 1-octen-3-ol (bOBP co-purified ligand) and other ligands with microM Kd values comparable to those of the swapped dimer.
- Dipartimento di Produzioni Animali, Biotecnologie Veterinarie, Qualità e Sicurezza degli Alimenti, Università degli Studi di Parma, Via del Taglio 8, 43100 Parma, Italy.
Organizational Affiliation: