Structural Determinants Involved in the Regulation of CXCL14/BRAK Expression by the 26 S Proteasome.
Peterson, F.C., Thorpe, J.A., Harder, A.G., Volkman, B.F., Schwarze, S.R.(2006) J Mol Biology 363: 813-822
- PubMed: 16987528
- DOI: https://doi.org/10.1016/j.jmb.2006.08.057
- Primary Citation of Related Structures:
2HDL - PubMed Abstract:
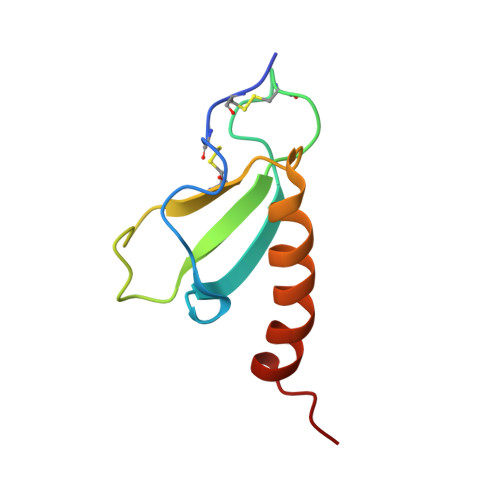
The chemokine CXCL14/BRAK participates in immune surveillance by recruiting dendritic cells. CXCL14 gene expression is altered in a number of cancers, but protein expression levels have not been investigated. Here we report that CXCL14 protein can be expressed in primary epithelial cells; however, in several immortalized and cancer cell lines this protein is targeted for polyubiquitylation and proteasomal degradation. We determined the NMR structure of CXCL14 to identify motifs controlling its expression. CXCL14 adopts the canonical chemokine tertiary fold but contains a unique five amino acid insertion (41VSRYR45) relative to other CXC chemokines. Deletion or substitution of key residues within this insertion prevented proteasomal degradation. Furthermore, we defined a 15 amino acid fragment of CXCL14 that is sufficient to induce proteasomal degradation. This study elucidates a post-translational mechanism for the loss of CXCL14 in cancer and a novel mode of chemokine regulation.
- Department of Biochemistry, Medical College of Wisconsin, Milwaukee, WI 53226, USA.
Organizational Affiliation: