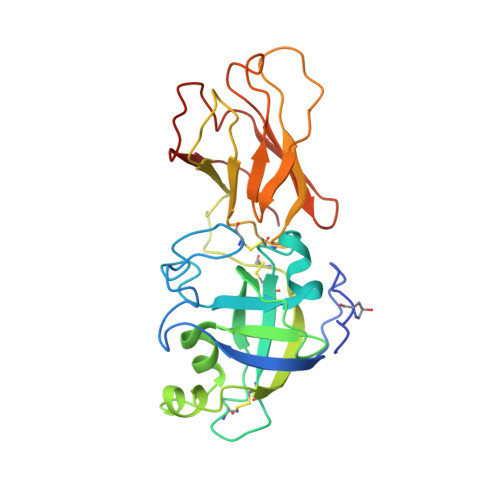
Crystal structure and activities of EXPB1 (Zea m 1), a beta-expansin and group-1 pollen allergen from maize.
Yennawar, N.H., Li, L.C., Dudzinski, D.M., Tabuchi, A., Cosgrove, D.J.(2006) Proc Natl Acad Sci U S A 103: 14664-14671
- PubMed: 16984999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0605979103
- Primary Citation Related Structures:
2HCZ - PubMed Abstract:
Expansins are small extracellular proteins that promote turgor-driven extension of plant cell walls. EXPB1 (also called Zea m 1) is a member of the beta-expansin subfamily known in the allergen literature as group-1 grass pollen allergens. EXPB1 induces extension and stress relaxation of grass cell walls. To help elucidate expansin's mechanism of wall loosening, we determined the structure of EXPB1 by x-ray crystallography to 2.75-A resolution. EXPB1 consists of two domains closely packed and aligned so as to form a long, shallow groove with potential to bind a glycan backbone of approximately 10 sugar residues. The structure of EXPB1 domain 1 resembles that of family-45 glycoside hydrolase (GH45), with conservation of most of the residues in the catalytic site. However, EXPB1 lacks a second aspartate that serves as the catalytic base required for hydrolytic activity in GH45 enzymes. Domain 2 of EXPB1 is an Ig-like beta-sandwich, with aromatic and polar residues that form a potential surface for polysaccharide binding in line with the glycan binding cleft of domain 1. EXPB1 binds to maize cell walls, most strongly to xylans, causing swelling of the cell wall. Tests for hydrolytic activity by EXPB1 with various wall polysaccharides proved negative. Moreover, GH45 enzymes and a GH45-related protein called "swollenin" lacked wall extension activity comparable to that of expansins. We propose a model of expansin action in which EXPB1 facilitates the local movement and stress relaxation of arabinoxylan-cellulose networks within the wall by noncovalent rearrangement of its target.
- Huck Institutes of the Life Sciences and Department of Biology, Pennsylvania State University, University Park, PA 16802, USA.
Organizational Affiliation: