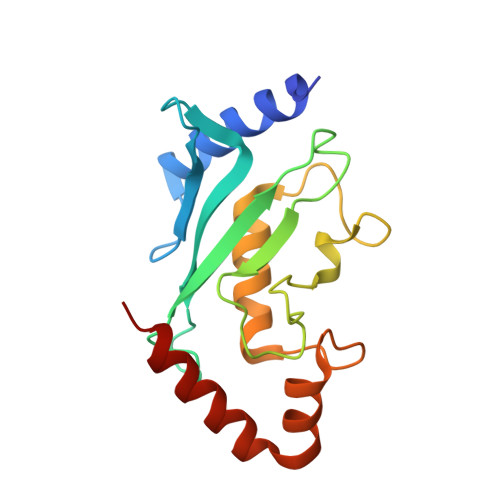
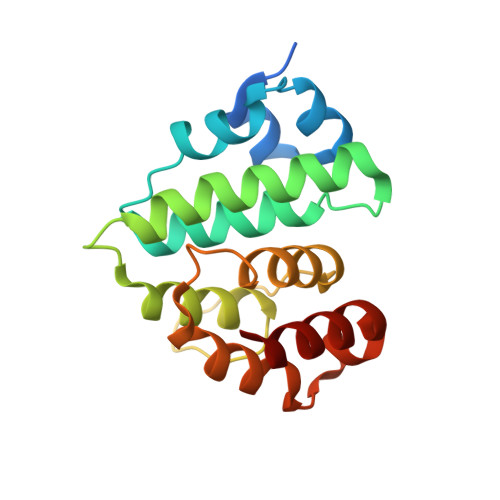
Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway.
Yunus, A.A., Lima, C.D.(2006) Nat Struct Mol Biol 13: 491-499
- PubMed: 16732283 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1104
- Primary Citation Related Structures:
2GRN, 2GRO, 2GRP, 2GRQ, 2GRR - PubMed Abstract:
E2 conjugating proteins that transfer ubiquitin and ubiquitin-like modifiers to substrate lysine residues must first activate the lysine nucleophile for conjugation. Genetic complementation revealed three side chains of the E2 Ubc9 that were crucial for normal growth. Kinetic analysis revealed modest binding defects but substantially lowered catalytic rates for these mutant alleles with respect to wild-type Ubc9. X-ray structures for wild-type and mutant human Ubc9-RanGAP1 complexes showed partial loss of contacts to the substrate lysine in mutant complexes. Computational analysis predicted pK perturbations for the substrate lysine, and Ubc9 mutations weakened pK suppression through improper side chain coordination. Biochemical studies with p53, RanGAP1 and the Nup358/RanBP2 E3 were used to determine rate constants and pK values, confirming both structural and computational predictions. It seems that Ubc9 uses an indirect mechanism to activate lysine for conjugation that may be conserved among E2 family members.
- Structural Biology Program, Sloan-Kettering Institute, New York, New York 10021, USA.
Organizational Affiliation: