Mis-translation of a Computationally Designed Protein Yields an Exceptionally Stable Homodimer: Implications for Protein Engineering and Evolution.
Dantas, G., Watters, A.L., Lunde, B.M., Eletr, Z.M., Isern, N.G., Roseman, T., Lipfert, J., Doniach, S., Tompa, M., Kuhlman, B., Stoddard, B.L., Varani, G., Baker, D.(2006) J Mol Biology 362: 1004-1024
- PubMed: 16949611 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.07.092
- Primary Citation Related Structures:
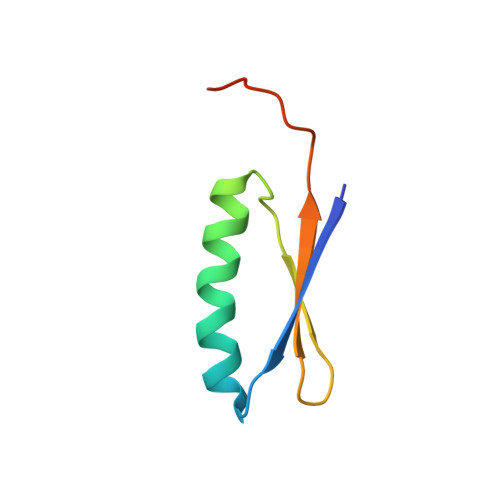
2GJH - PubMed Abstract:
We recently used computational protein design to create an extremely stable, globular protein, Top7, with a sequence and fold not observed previously in nature. Since Top7 was created in the absence of genetic selection, it provides a rare opportunity to investigate aspects of the cellular protein production and surveillance machinery that are subject to natural selection. Here we show that a portion of the Top7 protein corresponding to the final 49 C-terminal residues is efficiently mis-translated and accumulates at high levels in Escherichia coli. We used circular dichroism, size-exclusion chromatography, small-angle X-ray scattering, analytical ultra-centrifugation, and NMR spectroscopy to show that the resulting C-terminal fragment (CFr) protein adopts a compact, extremely stable, homo-dimeric structure. Based on the solution structure, we engineered an even more stable variant of CFr by disulfide-induced covalent circularisation that should be an excellent platform for design of novel functions. The accumulation of high levels of CFr exposes the high error rate of the protein translation machinery. The rarity of correspondingly stable fragments in natural proteins coupled with the observation that high quality ribosome binding sites are found to occur within E. coli protein-coding regions significantly less often than expected by random chance implies a stringent evolutionary pressure against protein sub-fragments that can independently fold into stable structures. The symmetric self-association between two identical mis-translated CFr sub-domains to generate an extremely stable structure parallels a mechanism for natural protein-fold evolution by modular recombination of protein sub-structures.
- Department of Biochemistry, University of Washington, Seattle 98195, USA.
Organizational Affiliation: