2-substituted estradiol bis-sulfamates, multitargeted antitumor agents: synthesis, in vitro SAR, protein crystallography, and in vivo activity.
Leese, M.P., Leblond, B., Smith, A., Newman, S.P., Di Fiore, A., De Simone, G., Supuran, C.T., Purohit, A., Reed, M.J., Potter, B.V.(2006) J Med Chem 49: 7683-7696
- PubMed: 17181151
- DOI: https://doi.org/10.1021/jm060705x
- Primary Citation of Related Structures:
2GD8 - PubMed Abstract:
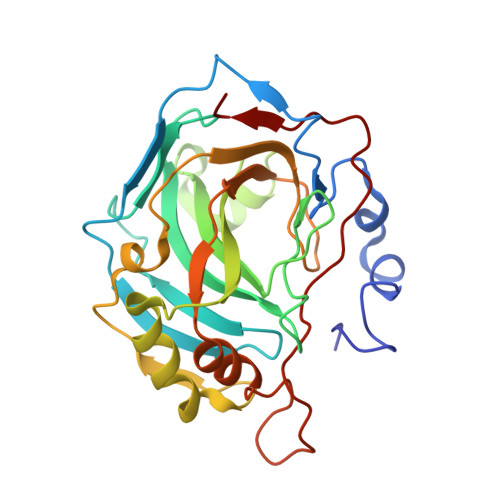
The anticancer activities and SARs of estradiol-17-O-sulfamates and estradiol 3,17-O,O-bis-sulfamates (E2bisMATEs) as steroid sulfatase (STS) inhibitors and antiproliferative agents are discussed. Estradiol 3,17-O,O-bis-sulfamates 20 and 21, in contrast to the 17-O-monosulfamate 11, proved to be excellent STS inhibitors. 2-Substituted E2bisMATEs 21 and 23 additionally exhibited potent antiproliferative activity with mean graph midpoint values of 18-87 nM in the NCI 60-cell-line panel. 21 Exhibited antiangiogenic in vitro and in vivo activity in an early-stage Lewis lung model, and 23 dosed p.o. caused marked growth inhibition in a nude mouse xenograft tumor model. Modeling studies suggest that the E2bisMATEs and 2-MeOE2 share a common mode of binding to tubulin, though COMPARE analysis of activity profiles was negative. 21 was cocrystallized with carbonic anhydrase II, and X-ray crystallography revealed unexpected coordination of the 17-O-sulfamate of 21 to the active site zinc and a probable additional lower affinity binding site. 2-Substituted E2bisMATEs are attractive candidates for further development as multitargeted anticancer agents.
- Medicinal Chemistry, Department of Pharmacy and Pharmacology & Sterix Ltd., University of Bath, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: