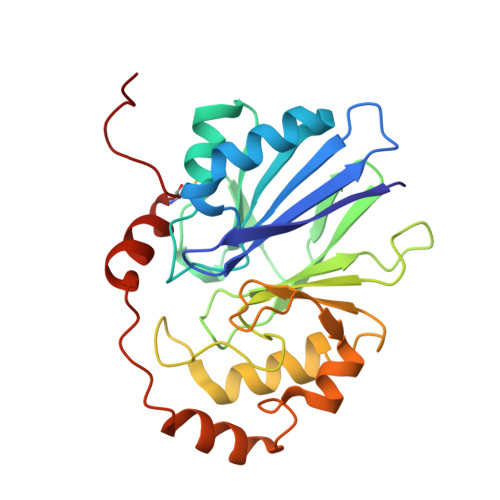
Structure of an ETHE1-like protein from Arabidopsis thaliana.
McCoy, J.G., Bingman, C.A., Bitto, E., Holdorf, M.M., Makaroff, C.A., Phillips, G.N.(2006) Acta Crystallogr D Biol Crystallogr 62: 964-970
- PubMed: 16929096 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444906020592
- Primary Citation Related Structures:
2GCU - PubMed Abstract:
The protein product of gene At1g53580 from Arabidopsis thaliana possesses 54% sequence identity to a human enzyme that has been implicated in the rare disorder ethylmalonic encephalopathy. The structure of the At1g53580 protein has been solved to a nominal resolution of 1.48 Angstrom. This structure reveals tertiary structure differences between the ETHE1-like enzyme and glyoxalase II enzymes that are likely to account for differences in reaction chemistry and multimeric state between the two types of enzymes. In addition, the Arabidopsis ETHE1 protein is used as a model to explain the significance of several mutations in the human enzyme that have been observed in patients with ethylmalonic encephalopathy.
- Department of Biochemistry and Center for Eukaryotic Structural Genomics, University of Wisconsin-Madison, WI, USA.
Organizational Affiliation: