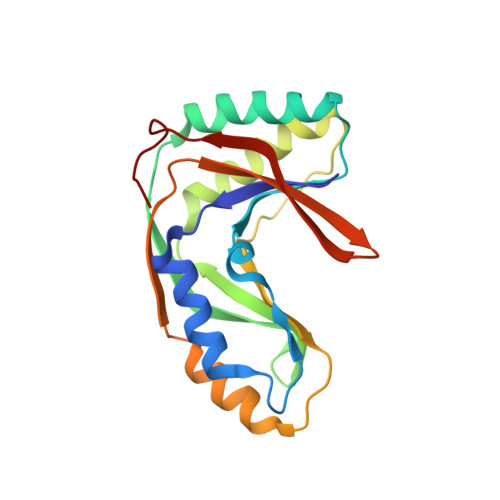
Characterization of a heat-stable enzyme possessing GTP-dependent RNA ligase activity from a hyperthermophilic archaeon, Pyrococcus furiosus
Kanai, A., Sato, A., Fukuda, Y., Okada, K., Matsuda, T., Sakamoto, T., Muto, Y., Yokoyama, S., Kawai, G., Tomita, M.(2009) RNA 15: 420-431
- PubMed: 19155324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.1122109
- Primary Citation Related Structures:
2FYH - PubMed Abstract:
Using an expression protein library of a hyperthermophilic archaeon, Pyrococcus furiosus, we identified a gene (PF0027) that encodes a protein with heat-stable cyclic nucleotide phosphodiesterase (CPDase) activity. The PF0027 gene encoded a 21-kDa protein and an amino acid sequence that showed approximately 27% identity to that of the 2'-5' tRNA ligase protein, ligT (20 kDa), from Escherichia coli. We found that the purified PF0027 protein possessed GTP-dependent RNA ligase activity and that synthetic tRNA halves bearing 2',3'-cyclic phosphate and 5'-OH termini were substrates for the ligation reaction in vitro. GTP hydrolysis was not required for the reaction, and GTPgammaS enhanced the tRNA ligation activity of PF0027 protein, suggesting that the ligation step is regulated by a novel mechanism. In comparison to the strong CPDase activity of the PF0027 protein, the RNA ligase activity itself was quite weak, and the ligation product was unstable during in vitro reaction. Finally, we used NMR to determine the solution structure of the PF0027 protein and discuss the implications of our results in understanding the role of the PF0027 protein.
- Institute for Advanced Biosciences, Keio University, Tsuruoka, Yamagata 997-0017, Japan. akio@sfc.keio.ac.jp
Organizational Affiliation: