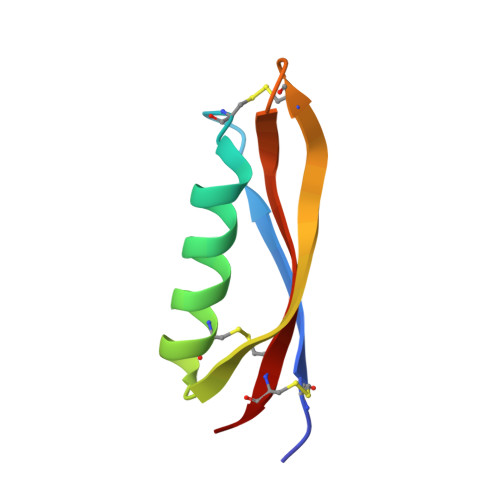
Structure of Alzheimer's disease amyloid precursor protein copper-binding domain at atomic resolution.
Kong, G.K., Adams, J.J., Cappai, R., Parker, M.W.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 819-824
- PubMed: 17909280
- DOI: https://doi.org/10.1107/S1744309107041139
- Primary Citation of Related Structures:
2FMA - PubMed Abstract:
Amyloid precursor protein (APP) plays a central role in the pathogenesis of Alzheimer's disease, as its cleavage generates the Abeta peptide that is toxic to cells. APP is able to bind Cu2+ and reduce it to Cu+ through its copper-binding domain (CuBD). The interaction between Cu2+ and APP leads to a decrease in Abeta production and to alleviation of the symptoms of the disease in mouse models. Structural studies of CuBD have been undertaken in order to better understand the mechanism behind the process. Here, the crystal structure of CuBD in the metal-free form determined to ultrahigh resolution (0.85 A) is reported. The structure shows that the copper-binding residues of CuBD are rather rigid but that Met170, which is thought to be the electron source for Cu2+ reduction, adopts two different side-chain conformations. These observations shed light on the copper-binding and redox mechanisms of CuBD. The structure of CuBD at atomic resolution provides an accurate framework for structure-based design of molecules that will deplete Abeta production.
- Biota Structural Biology Laboratory, St Vincent's Institute, 9 Princes Street, Fitzroy, Victoria 3065, Australia.
Organizational Affiliation: