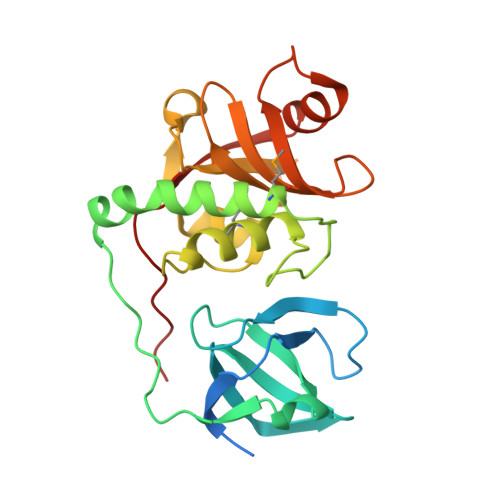
Structural Basis of Murein Peptide Specificity of a gamma-D-Glutamyl-L-Diamino Acid Endopeptidase.
Xu, Q., Sudek, S., McMullan, D., Miller, M.D., Geierstanger, B., Jones, D.H., Krishna, S.S., Spraggon, G., Bursalay, B., Abdubek, P., Acosta, C., Ambing, E., Astakhova, T., Axelrod, H.L., Carlton, D., Caruthers, J., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Elias, Y., Elsliger, M.A., Feuerhelm, J., Grzechnik, S.K., Hale, J., Won Han, G., Haugen, J., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Kumar, A., Marciano, D., Morse, A.T., Nigoghossian, E., Okach, L., Oommachen, S., Paulsen, J., Reyes, R., Rife, C.L., Trout, C.V., van den Bedem, H., Weekes, D., White, A., Wolf, G., Zubieta, C., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2009) Structure 17: 303-313
- PubMed: 19217401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.12.008
- Primary Citation Related Structures:
2EVR, 2FG0, 2HBW - PubMed Abstract:
The crystal structures of two homologous endopeptidases from cyanobacteria Anabaena variabilis and Nostoc punctiforme were determined at 1.05 and 1.60 A resolution, respectively, and contain a bacterial SH3-like domain (SH3b) and a ubiquitous cell-wall-associated NlpC/P60 (or CHAP) cysteine peptidase domain. The NlpC/P60 domain is a primitive, papain-like peptidase in the CA clan of cysteine peptidases with a Cys126/His176/His188 catalytic triad and a conserved catalytic core. We deduced from structure and sequence analysis, and then experimentally, that these two proteins act as gamma-D-glutamyl-L-diamino acid endopeptidases (EC 3.4.22.-). The active site is located near the interface between the SH3b and NlpC/P60 domains, where the SH3b domain may help define substrate specificity, instead of functioning as a targeting domain, so that only muropeptides with an N-terminal L-alanine can bind to the active site.
- Joint Center for Structural Genomics, SLAC National Accelerator Laboratory, Stanford University, Menlo Park, CA 94025, USA.
Organizational Affiliation: