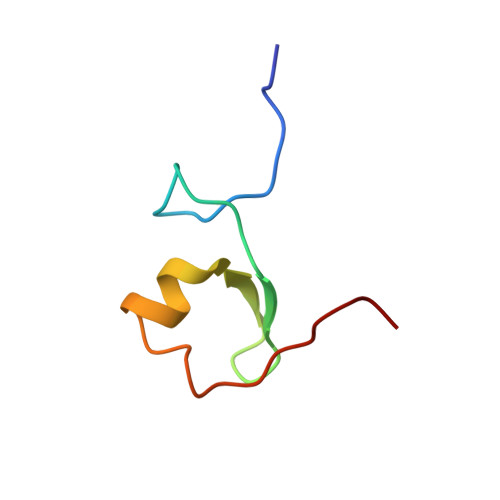
Solution Structure of the MID1 B-box2 CHC(D/C)C(2)H(2) Zinc-binding Domain: Insights into an Evolutionarily Conserved RING Fold
Massiah, M.A., Matts, J.A.B., Short, K.M., Simmons, B.N., Singireddy, S., Yi, Z., Cox, T.C.(2007) J Mol Biology 369: 1-10
- PubMed: 17428496 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.03.017
- Primary Citation Related Structures:
2DQ5 - PubMed Abstract:
The B-box type 2 domain is a prominent feature of a large and growing family of RING, B-box, coiled-coil (RBCC) domain-containing proteins and is also present in more than 1500 additional proteins. Most proteins usually contain a single B-box2 domain, although some proteins contain tandem domains consisting of both type 1 and type 2 B-boxes, which actually share little sequence similarity. Recently, we determined the solution structure of B-box1 from MID1, a putative E3 ubiquitin ligase that is mutated in X-linked Opitz G/BBB syndrome, and showed that it adopted a betabetaalpha RING-like fold. Here, we report the tertiary structure of the B-box2 (CHC(D/C)C(2)H(2)) domain from MID1 using multidimensional NMR spectroscopy. This MID1 B-box2 domain consists of a short alpha-helix and a structured loop with two short anti-parallel beta-strands and adopts a tertiary structure similar to the B-box1 and RING structures, even though there is minimal primary sequence similarity between these domains. By mutagenesis, ESI-FTICR and ICP mass spectrometry, we show that the B-box2 domain coordinates two zinc atoms with a 'cross-brace' pattern: one by Cys175, His178, Cys195 and Cys198 and the other by Cys187, Asp190, His204, and His207. Interestingly, this is the first case that an aspartic acid is involved in zinc atom coordination in a zinc-finger domain, although aspartic acid has been shown to coordinate non-catalytic zinc in matrix metalloproteinases. In addition, the finding of a Cys195Phe substitution identified in a patient with X-linked Opitz GBBB syndrome supports the importance of proper zinc coordination for the function of the MID1 B-box2 domain. Notably, however, our structure differs from the only other published B-box2 structure, that from XNF7, which was shown to coordinate one zinc atom. Finally, the similarity in tertiary structures of the B-box2, B-box1 and RING domains suggests these domains have evolved from a common ancestor.
- Department of Biochemistry and Molecular Biology, Oklahoma State University, Stillwater, OK 74078, USA. massiah@biochem.okstate.edu
Organizational Affiliation: