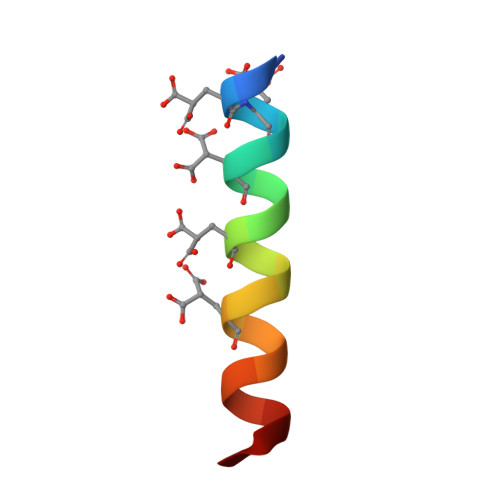
The crystal structures of the calcium-bound con-G and con-T[K7gamma] dimeric peptides demonstrate a metal-dependent helix-forming motif
Cnudde, S.E., Prorok, M., Dai, Q., Castellino, F.J., Geiger, J.H.(2007) J Am Chem Soc 129: 1586-1593
- PubMed: 17243678 Search on PubMed
- DOI: https://doi.org/10.1021/ja065722q
- Primary Citation Related Structures:
2DPQ, 2DPR - PubMed Abstract:
Short peptides that have the ability to form stable alpha-helices in solution are rare, and a number of strategies have been used to produce them, including the use of metal chelation to stabilize folding of the backbone. However, no example exists of a structurally well-defined helix stabilized exclusively through metal ion chelation. Conantokins (con)-G and -T are short peptides that are potent antagonists of N-methyl-D-aspartate receptor channels. While con-G exhibits no helicity alone, it undergoes a structural transition to a helical conformation in the presence of a variety of multivalent cations, especially Mg2+ and Ca2+. This complexation also results in antiparallel dimerization of two peptide helices in the presence of Ca2+, but not Mg2+. A con-T variant, con-T[K7gamma], displays very similar behavior. We have solved the crystal structures of both Ca2+/con-G and Ca2+/con-T [K7gamma] at atomic resolution. These structures clearly show the nature of the metal-dependent dimerization and helix formation and surprisingly also show that the con-G dimer interface is completely different from the con-T[K7gamma] interface, even though the metal chelation is similar in the two peptides. This represents a new paradigm in helix stabilization completely independent of the hydrophobic effect, which we define as the "metallo-zipper."
- Department of Biochemistry, Michigan State University, East Lansing, Michigan 48824, USA.
Organizational Affiliation: