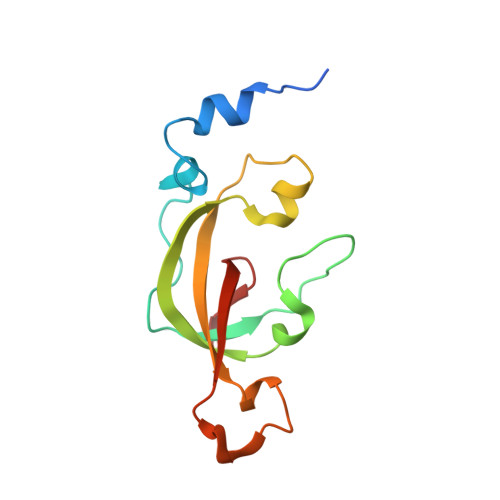
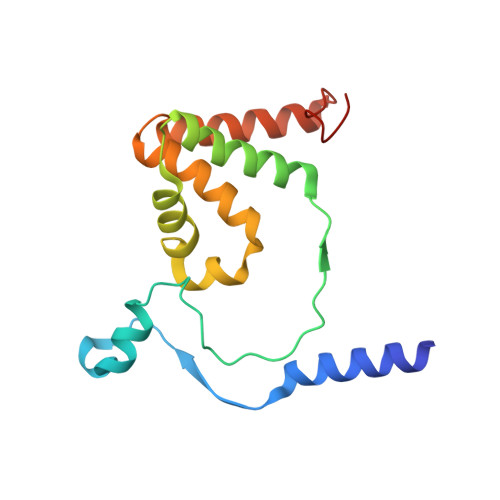
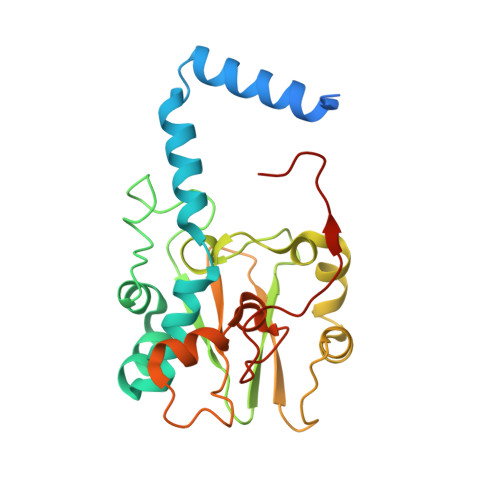
Structure of thiocyanate hydrolase: a new nitrile hydratase family protein with a novel five-coordinate cobalt(III) center.
Arakawa, T., Kawano, Y., Kataoka, S., Katayama, Y., Kamiya, N., Yohda, M., Odaka, M.(2007) J Mol Biology 366: 1497-1509
- PubMed: 17222425
- DOI: https://doi.org/10.1016/j.jmb.2006.12.011
- Primary Citation Related Structures:
2DD4, 2DD5 - PubMed Abstract:
Thiocyanate hydrolase (SCNase) of Thiobacillus thioparus THI115 is a cobalt(III)-containing enzyme catalyzing the degradation of thiocyanate to carbonyl sulfide and ammonia. We determined the crystal structures of the apo- and native SCNases at a resolution of 2.0 A. SCNases in both forms had a conserved hetero-dodecameric structure, (alphabetagamma)(4). Four alphabetagamma hetero-trimers were structurally equivalent. One alphabetagamma hetero-trimer was composed of the core domain and the betaN domain, which was located at the center of the molecule and linked the hetero-trimers with novel quaternary interfaces. In both the apo- and native SCNases, the core domain was structurally conserved between those of iron and cobalt-types of nitrile hydratase (NHase). Native SCNase possessed the post-translationally modified cysteine ligands, gammaCys131-SO(2)H and gammaCys133-SOH like NHases. However, the low-spin cobalt(III) was found to be in the distorted square-pyramidal geometry, which had not been reported before in any protein. The size as well as the electrostatic properties of the substrate-binding pocket was totally different from NHases with respect to the charge distribution and the substrate accessibility, which rationally explains the differences in the substrate preference between SCNase and NHase.
- Department of Biotechnology and Life Science, Graduate School of Technology, Tokyo University of Agriculture and Technology, 2-24-16 Naka-cho, Koganei, Tokyo 184-8588, Japan.
Organizational Affiliation: