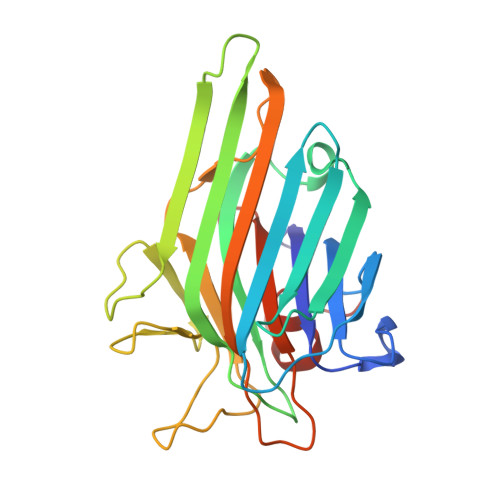
Native crystal structure of a nitric oxide-releasing lectin from the seeds of Canavalia maritima
Gadelha, C.A.A., Moreno, F.B.M.B., Santi-Gadelha, T., Cajazeiras, J.B., Rocha, B.A.M., Assreuy, A.M.S., Lima Mota, M.R., Pinto, N.V., Passos Meireles, A.V., Borges, J.C., Freitas, B.T., Canduri, F., Souza, E.P., Delatorre, P., Criddle, D.N., De Azevedo Jr., W.F., Cavada, B.S.(2005) J Struct Biol 152: 185-194
- PubMed: 16337811 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2005.07.012
- Primary Citation Related Structures:
2CWM - PubMed Abstract:
Here, we report the crystallographic study of a lectin from Canavalia maritima seeds (ConM) and its relaxant activity on vascular smooth muscle, to provide new insights into the understanding of structure/function relationships of this class of proteins. ConM was crystallized and its structure determined by standard molecular replacement techniques. The amino acid residues, previously suggested incorrectly by manual sequencing, have now been determined as I17, I53, S129, S134, G144, S164, P165, S187, V190, S169, T196, and S202. Analysis of the structure indicated a dimer in the asymmetric unit, two metal binding sites per monomer, and loops involved in the molecular oligomerization. These confer 98% similarity between ConM and other previously described lectins, derived from Canavalia ensiformis and Canavalia brasiliensis. Our functional data indicate that ConM exerts a concentration-dependent relaxant action on isolated aortic rings that probably occurs via an interaction with a specific lectin-binding site on the endothelium, resulting in a release of nitric oxide.
- BioMol-Lab/Departamento de Bioquímica e Biologia Molecular, Universidade Federal do Ceará, Brazil.
Organizational Affiliation: