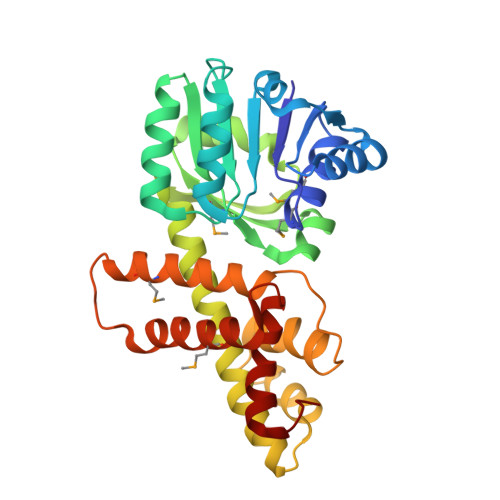
Crystal Structure of Novel NADP-dependent 3-Hydroxyisobutyrate Dehydrogenase from Thermus thermophilus HB8
Lokanath, N.K., Ohshima, N., Takio, K., Shiromizu, I., Kuroishi, C., Okazaki, N., Kuramitsu, S., Yokoyama, S., Miyano, M., Kunishima, N.(2005) J Mol Biology 352: 905-917
- PubMed: 16126223 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.068
- Primary Citation Related Structures:
1WP4, 2CVZ - PubMed Abstract:
3-Hydroxyisobutyrate, a central metabolite in the valine catabolic pathway, is reversibly oxidized to methylmalonate semialdehyde by a specific dehydrogenase belonging to the 3-hydroxyacid dehydrogenase family. To gain insight into the function of this enzyme at the atomic level, we have determined the first crystal structures of the 3-hydroxyisobutyrate dehydrogenase from Thermus thermophilus HB8: holo enzyme and sulfate ion complex. The crystal structures reveal a unique tetrameric oligomerization and a bound cofactor NADP+. This bacterial enzyme may adopt a novel cofactor-dependence on NADP, whereas NAD is preferred in eukaryotic enzymes. The protomer folds into two distinct domains with open/closed interdomain conformations. The cofactor NADP+ with syn nicotinamide and the sulfate ion are bound to distinct sites located at the interdomain cleft of the protomer through an induced-fit domain closure upon cofactor binding. From the structural comparison with the crystal structure of 6-phosphogluconate dehydrogenase, another member of the 3-hydroxyacid dehydrogenase family, it is suggested that the observed sulfate ion and the substrate 3-hydroxyisobutyrate share the same binding pocket. The observed oligomeric state might be important for the catalytic function through forming the active site involving two adjacent subunits, which seems to be conserved in the 3-hydroxyacid dehydrogenases. A kinetic study confirms that this enzyme has strict substrate specificity for 3-hydroxyisobutyrate and serine, but it cannot distinguish the chirality of the substrates. Lys165 is likely the catalytic residue of the enzyme.
- Highthroughput Factory, RIKEN Harima Institute at SPring-8, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: