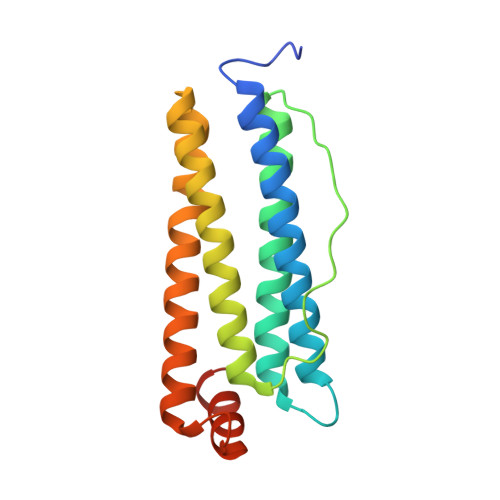
High-Resolution X-Ray Structures of Human Apoferritin H-Chain Mutants Correlated with Their Activity and Metal-Binding Sites.
Toussaint, L., Bertrand, L., Hue, L., Crichton, R.R., Declercq, J.P.(2007) J Mol Biology 365: 440
- PubMed: 17070541 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.10.010
- Primary Citation Related Structures:
2CEI, 2CHI, 2CIH, 2CLU, 2CN6, 2CN7, 2IU2 - PubMed Abstract:
Ferritins are a family of proteins distributed widely in nature. In bacterial, plant, and animal cells, ferritin appears to serve as a soluble, bioavailable, and non-toxic form of iron provider. Ferritins from animal sources are heteropolymers composed of two types of subunit, H and L, which differ mainly by the presence (H) or absence (L) of active ferroxidase centres. We report the crystallographic structures of four human H apoferritin variants at a resolution of up to 1.5 Angstrom. Crystal derivatives using Zn(II) as redox-stable alternative for Fe(II), allows us to characterize the different metal-binding sites. The ferroxidase centre, which is composed of sites A and B, binds metal with a preference for the A site. In addition, distinct Zn(II)-binding sites were found in the 3-fold axes, 4-fold axes and on the cavity surface near the ferroxidase centre. To study the importance of the distance of the two metal atoms in the ferroxidase centre, single and double replacement of glutamate 27 (site A) and glutamate 107 (site B), the two axial ligands, by aspartate residues have been carried out. The consequences for metal binding and the correlation with Fe(II) oxidation rates are discussed.
- Biochemistry Unit, Université Catholique de Louvain, 1 Place Louis Pasteur, 1348 Louvain la Neuve, Belgium.
Organizational Affiliation: